Figure 4.
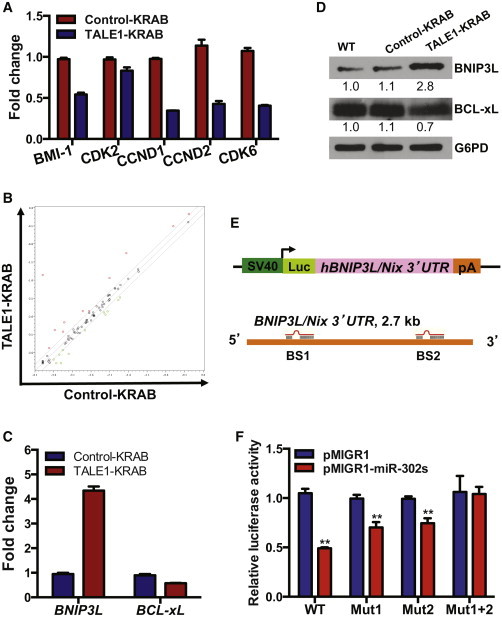
Target Genes of miR-302/367 Cluster Associated with Cell Cycle and Apoptosis in hESCs
(A) qPCR analysis of potential targets of miR-302/367 cluster regulating cell cycle in hESCs. Data are represented as mean ± SD of technical replicates (n = 3). See also Figure S2 and Table S1.
(B) Screening of apoptosis-related target genes of the miR-302/367 cluster. Patterns of gene expression were compared between the hESCs expressing control-KRAB or TALE1-KRAB using Apoptosis PCR Array. See also Table S2.
(C) qPCR analysis of BNIP3L and BCL-xL gene expression in control-KRAB- and TALE1-KRAB-expressing hESCs. Data are represented as mean ± SD of technical replicates (n = 3).
(D) Western blot analysis of BINP3L/Nix and BCL-xL in control-KRAB- and TALE1-KRAB-expressing hESCs. BINP3L/Nix and BCL-xL were detected by western blot analysis with each specific antibody. Relative expression of the BINP3L/Nix and BCL-xL was quantified by Image J (NIH) and normalized by G6PD.
(E) Schematic representation of the reporter construct containing the luciferase-coding sequence fused to the BNIP3L/Nix 3′UTR. Two predicted targeting sites for miR-302/367 cluster were designated as BS1 and BS2. See also Figure S3.
(F) BNIP3L/Nix 3′UTR luciferase reporter assay. 293T cells were transfected with the reporter plasmid pGL3-BNIP3L/Nix carrying mutations BS1 or BS2 or both, together with pMIGR1 (vector control) or pMIGR1_miR-302/367 and then cultured for 72 hr before luciferase activity assay. pCMV-LacZ was included in each transfection as an internal control to normalize luciferase activity. Data are represented as mean ± SD of three independent experiments (∗∗p < 0.01).
