Abstract
Chromophores that incorporate f-block elements have considerable potential for use in bioimaging applications because of their advantageous photophysical properties compared to organic dye, which are currently widely used. We are developing new classes of lanthanide-based self-assembling molecular nanoparticles as reporters for imaging and as multi-functional nanoprobes or nanosensors for use with biological samples. One class of these materials, which we call lanthanide “nano-drums”, are homogeneous 4d–4f clusters approximately 25 to 30 Å in diameter. These are capable of emitting from the visible to near-infrared wavelengths. Here, we present the synthesis, crystal structure, photophysical properties and comparative cytotoxicity data for a 32 metal Eu-Cd nano-drum [Eu8Cd24L12(OAc)48] (1). We also explored the imaging capabilities of this nano-drum using epifluorescence, TIRF, and two-photon microscopy platforms.
1. Introduction
Many imaging and diagnostics applications rely upon the use of fluorescent dyes or labels as reporter molecules because of their exceptional measurement contrast properties.1 The bulk of compounds in use as reporter molecules are organic dyes or genetically-encoded fluorescent proteins. New multiphoton imaging technologies now enable experiments to be performed within the complex environment of living tissues, enabling real-time analyses of cell function and structure to be undertaken.2 However, it is well-recognized that both fluorescent dyes and protein-based fluorescent probes have technical limitations including susceptibility to severe photo-bleaching (which limits long-term analysis) and small Stokes shifts (which limit the utility of the emission range). Development of new probes or labels with improved photophysical properties is clearly desirable, particularly for use in techniques such as multiphoton microscopy.2
The use of lanthanide ions (Ln3+) as photoluminescent bioprobes in either small molecules3,4 or nanocrystals of a host matrix comprising oxides,5–7 vanadates,8,9 oxysulfides,10,11 phosphates12,13 or fluorides14–18 is well known. Very recently an interesting new system was reported in which 3-dimensional network structures of lanthanide ions and organic linkers (metal–organic frameworks or MOFs) were used for imaging in the near infra-red in living cells.19 The advantages of Ln3+ based bioprobes over conventional biomarkers based on organic dyes or semiconductor quantum dots are also well documented. For example, conventional bioprobes can suffer from high autofluorescence background and photodamage associated with ultraviolet excitation, large emission bandwidths, low photochemical stability, short luminescent lifetimes and long-term toxicity. In contrast, lanthanide based bioprobes exhibit a very large pseudo-Stokes shift between the excitation and emission wavelengths, absence of photo-bleaching, long lived excited states and narrow emission bands which enable selective detection.3,4,20,21 Since emissions via 4f–4f transitions are parity forbidden,22 Ln3+ ions usually display relatively long photoluminescence lifetimes following photoexcitation. This property permits gated and time resolved photoluminescence detection thus eliminating interference from short-lived autofluorescence. In molecular compounds the weak 4f–4f emissions can be sensitized by suitable ligand chromophores, which transfer energy from the triplet state of the ligand to the Ln3+ ion. This “antenna” effect can result in strong and useful luminescence, which is observed in the visible region for Eu3+, Tb3+, Sm3+ and Dy3+.23,24
As part of our research focused on high nuclearity lanthanide moieties we recently reported a new class of luminescent high nuclearity self-assembled cadmium-lanthanide cluster molecules (“molecular nanoparticles”) with unusual drum-like architectures.25,26 Our original report described nano-drum near-infrared (NIR) emitters that featured Nd, Yb, Er, as well as Gd.26 Since many lanthanide-based probes systems employ Eu3+ which emits in the visible spectrum, we have investigated the synthesis and photophysical properties of the 32-metal molecular nano-drum based on this element: [Eu8Cd24L12(OAc)48] (1). The complex has been structurally characterized by single crystal X-ray diffraction studies and tunnelling electron microscopy. Its photophysical properties were also investigated. We report initial studies aimed at evaluating the potential of 1 to serve as a dual action cytotoxic fluorescent probe in biological environments. We have found that the nano-drum 1 displays good antiproliferative activity, displaying an IC50 value of 1.37 ± 0.43 µM and 2.0 µM in an A549 lung cancer cell line and an AGS gastric cancer cell line, respectively. The imaging properties of 1 were investigated using three microscopy platforms commonly used for visualization of cells and tissues: epifluorescence, total internal reflection fluorescence (TIRF), and two-photon microscopy. On the basis of the present findings, we propose that complex 1 may have utility in a number of biological applications.
2. Experimental
2.1 Synthesis of [Eu8Cd24L12(OAc)48] (1)
All reactions were performed under dry oxygen-free dinitrogen atmospheres using standard Schlenk techniques. Metal salts and other solvents were purchased from Aldrich and used directly without further purification. The Schiff-base ligand H2L (N,N′-bis(3-methoxysalicylidene)hexane-1,6-diamine) was prepared according to well-established procedures.27 Melting points were obtained in sealed glass capillaries under dinitrogen and are uncorrected.
[Eu8Cd24L12(OAc)48] (1)
Eu(OAc)3·4H2O, (0.0501 g, 0.12 mmol), Cd(OAc)2·2H2O, (0.1384 g, 0.52 mmol) and H2L (0.1000 g, 0.26 mmol) were dissolved in MeOH (25 mL) at room temperature and Et3N (1 mL) added. The resulting solution was stirred and heated under reflux for 30 min. After cooling to room temperature, the solution was filtered and the filtrate transferred to test tubes. Diethyl ether vapor was allowed to diffuse slowly into the filtrate at room temperature and pale yellow crystals were obtained after one week. The crystals were filtered off and washed with MeOH (2 ml). Yield (based on Eu(OAc)3·4H2O): 0.0880 g (52%). m. p. >178 °C (dec.). IR (ATR, cm−1): 2932 (w), 2855 (w), 1889 (w), 1632 (m), 1584 (s), 1467 (s), 1345 (w), 1304 (m), 1237 (w), 1210 (s), 1168 (w), 1096 (w), 1078 (m), 1047 (w), 1015 (m), 961 (m), 853 (m), 782 (w), 737 (s), 666 (m), 640 (w), 613 (m), 551 (w), 494 (w), 455 (w), 435 (w), 417 (w).
2.2 X-ray structure determination
Powder X-ray diffraction data were collected on a Rigaku R-AXIS RAPID II. Single crystal data were collected on a Rigaku Saturn Kappa CCD diffractometer with graphite monochromated Mo-Kα radiation (λ = 0.71073 Å) at 223 K. The data set was corrected for absorption based on multiple scans and reduced using standard methods. Data reduction was performed using DENZO-SMN.28 The structure was solved by direct methods and refined anisotropically using full-matrix least-squares methods with the SHELX 97 program package.29 Coordinates of the non-hydrogen atoms were refined anisotropically, while hydrogen atoms were included in the calculation isotropically but not refined. Neutral atom scattering factors were taken from Cromer and Waber.30 Crystallographic data is given in the ESI (Tables S1 and S2†). (CCDC reference number 1008697. See http://www.rsc.org/suppdata/cc for crystallographic data in CIF format).
2.3 Bead preparation
Polystyrene beads (6 µm, crosslinked) were suspended in a 50 : 50 (v/v) solution of methanol and chloroform in 1.5 mL microfuge tubes. Typically 0.5–1 mg of 1 was added and the mixture was then rotated from 2–4 days to permit bead loading. Suspensions were then dried under vacuum overnight and beads removed and then washed and re-suspended in phosphate buffered saline (PBS) prior to imaging.
2.4 Spectroscopic characterization
Physical measurements were made using the following instrumentation: NMR: VARIAN UNITY-plus 600 spectrometer (1H, 600 MHz) at 298 K in CD3OD; IR: Nicolet IR 200 FTIR spectrometer. Absorption spectra were obtained on a BECKMAN DU 640 spectrophotometer, excitation and visible emission spectra on a QuantaMaster PTI fluorimeter. Fluorescence quantum yields were determined by using [Ru(bipy)3]Cl2 (bipy = 2,2′-bipyridine;Φem = 0.028 in water) as standard for the Eu3+ complex.31
X-ray photoelectron spectra (XPS) were obtained using a Kratos AXIS Ultra spectrometer equipped with a monochromatized Al Kα source, hybrid optics, and a delay line detector coupled to a hemispherical analyzer. The analysis chamber base pressure was typically 2 × 10−9 Torr. All spectra were recorded using a single sweep with a spot size of 300 µm × 700 µm. Survey scans were collected from 0–1200 eV with a pass energy of 80 eV, step size of 1 eV, and a dwell time of 250 ms. High resolution component spectra were collected with a pass energy of 20 eV, step size of 0.1 eV, and a dwell time of 3600 ms. Binding energies were referenced to the adventitious carbon line (C 1s, 284.8 eV) and a charge neutralization was applied during all acquisitions. Casa XPS analysis software was used for stoichiometry determination of the samples and Kratos sensitivity factors used for each element of interest. All samples were washed with methanol three times and dried under vacuum for 18 h before placing in an Al cup within the XPS instrument. Samples were carefully spread evenly within the cup using a blunt tool covered with Al foil between each procedure to avoid contamination.
2.5 Cytotoxicity assays
The proliferation of exponential phase cultures were carried out using either an A549 or an AGS cancer cell line. A549 cells were seeded in 96-well microliter plates at 1000 cells per well and allowed to adhere overnight in 100 µL RPMI 1640 medium supplemented with 2 mM l-glutamine, 10% heat inactivated fetal bovine serum, and antibiotics (200 U cm−3 penicillin and 200 µg/cm−3 streptomycin). AGS cells were seeded in 96-well microliter plates at 1000, 2000 and 4000 cells per well, grown in F-12 medium supplemented with 10% heat inactivated fetal bovine serum, and 100 U ml−1 penicillin and 100 µg ml−1 streptomycin, and allowed to adhere for 1–2 days. Cell viability was assessed by tetrazolium salt reduction. Stock solutions of [Eu8Cd24L12(OAc)48] (1), Eu(OAc)3·4H2O, Cd(OAc)2·2H2O, and H2L (all 5 mM) in 50/50 methanol–water were formulated and then diluted in medium for secondary stocks of 20–200 µM depending on the complex being tested. Secondary stock solutions were serially diluted in medium and immediately added to wells, whereupon plates were incubated at 37 °C under a 5% CO2/95% air atmosphere. After a total of 3 days, a 50 µL aliquot of 3 mg mL−1 tetrazolium dye, 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT, Sigma Chemical), was added to each well, followed by incubation for 4 h at 37 °C. The medium was then removed and the resulting formazan was dissolved in 50 µL DMSO and absorbances measured at 560–650 nm using a microplate reader (Molecular Devices, Sunnyvale, CA). Absorbances were corrected for background and the values normalized to wells containing untreated cells to allow plate-to-plate comparison. The data are shown as mean inhibition of proliferation or growth corresponding to the percentage of control cells’ proliferation or growth from 2–3 replicate values.
2.6 Microscopy
A flow chamber was constructed from a precleaned glass slide and was coverslip separated by use of a double-sided sticky tape. About 1 mg of polystyrene beads, loaded with either 1 or H2L ligand loaded beads were suspended in 20 µL of water and loaded in the flow chamber, which was used for both the Epifluorescent and TIRF imaging. For epifluorescence imaging, the slide was mounted on a Nikon Eclipse TE2000-E inverted microscope (Nikon Inc, Japan) and images of the beads acquired at 1 frame s−1 using a Cascade II 512 camera (Photometrics, Tucson, AZ) on a Nikon Apo 60×/NA 0.95 objective. A combination of excitation filters DAPI – AT350/50× (340–380 nm; Center Wavelength 360 nm), FITC – ET490/20× (465–495 nm; Center Wavelength 490 nm) or TRITC – ET555/25× (528–553 nm; Center Wavelength 555 nm) and emission filters DAPI – ET460/50 m (435–485 nm; Center Wavelength 460 nm), FITC – ET525/36 m (515–555 nm; Center wavelength 525 nm), TRITC – ET605/52 m (590–650 nm; Center wavelength 605 nm) or Cy5 – ET700/60× (640–730 nm; Center wavelength 700 nm) were used to image the lanthanide dye. The Sutter Lambda 10-3 filter wheels (Sutter Instrument, Novato, CA), the motorized stage (Prior Scientific Inc; Rockland, MA), and the image analysis were driven by Nikon NIS Elements Imaging Software.
Total internal reflection fluorescence (TIRF) imaging was performed in the objective mode using a Nikon Apo 60×/NA 1.49 oil objective set on a Nikon Eclipse Ti inverted microscope, fitted with a ProScan II motorized stage (Prior Scientific Inc; Rockland, MA) and a monolithic laser launch unit (Agilent Technologies; Santa Clara, CA) that is capable of providing tunable power and wavelength. Beads were excited by the 405 nm laser at 2 mW power (at source of fiber optic) and images acquired using an EMCCD iXon 3 camera (Andor; Belfast, UK) at 1 frame s−1 exposures before being filtered by a TIRF Quad Cube (Chroma Technology, Bellows Falls, VT). The Quad band filter is a configuration of BandPass filters and dichroics that allow for 405/488/561/647 ± 10 nm excitation and emission between 417–476 nm, 500–547 nm, 574–620 nm and 637–670 nm.
Two-photon fluorescence microscopy images were acquired with an Ultima IV microscope using PrairieView acquisition software and an Olympus Plan FL 20× water immersion objective (Prairie Technologies; Middleton, WI). Beads were illuminated with a MaiTai titanium:sapphire laser (Spectral Physics; Santa Clara, CA) tuned to 760 nm, or scanned from 690–1000 nm in 10 nm steps. Emitted light was passed through 400/50 nm, 480/40 nm, 535/50 nm, and 607/45 nm bandpass filters (Chroma Technology; Bellows Falls, VT) to individual PMTs for detection.
Transmission electron microscopy (TEM) images were recorded on a JEOL JEM-1200EX transmission electron microscope. Field emission scanning electron microscopy (FESEM) images were recorded on a Nova NanoSEM 200 scanning electron microscope.
3. Results
3.1 Synthesis and X-ray structure of nano-drum [Eu8Cd24L12(OAc)48] (1)
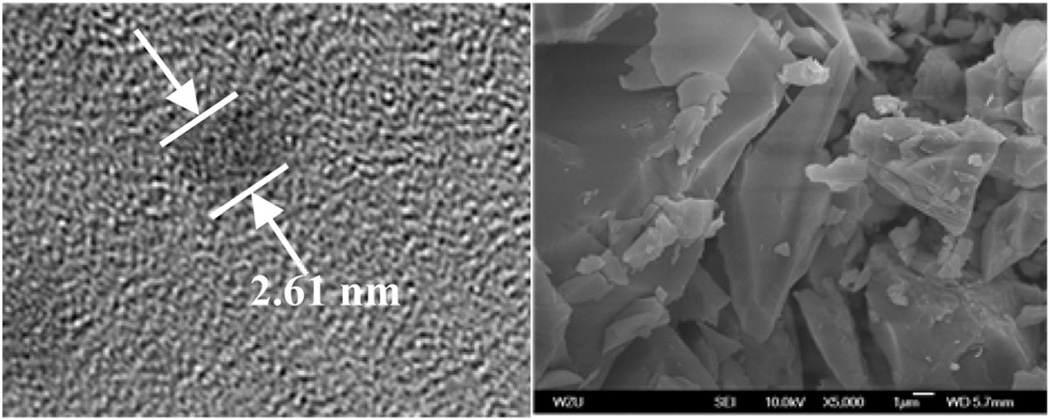
Reaction of the flexible Schiff base ligand H2L (Fig. 1) with Cd(OAc)2·4H2O and Eu(OAc)3·4H2O in MeOH–EtOH at reflux produced yellow solutions from which the nano-drum complex [Eu8Cd24L12(OAc)48] (1) was isolated as a pale yellow crystalline solid in approximately 50% yield. A key factor affecting self-assembly process is the use of the flexible Schiff-base ligand H2L which facilitates the construction of the nano-drum. The central six carbon –(CH2)6– chains of the Schiff base ligands form the sides of the drum while the deprotonated phenolic oxygen atoms, imine nitrogen and methoxy oxygen atoms bind the metals at the top and bottom of the drum. Views of the crystal structure of 1 are shown in Fig. 2 along with a space filling diagram. The top and bottom of the drum are created by two rings of 16 metals (4 Eu(III) and 12 Cd(II)) coordinated to half of the N- and O-binding groups of the 12 Schiff base ligands, plus 24 OAc− anions. In each 16-metal ring, four Eu ions are eight-coordinate. Of the twelve Cd2+ ions, four have bi-pyramidal geometries, and the other eight are seven-coordinate. The X-ray crystallographic data reveal that the molecular dimensions of 1 are approximately 24 Å × 26 Å × 26 Å. By virtue of the nanoscale dimensions of 1, it is also possible to obtain images of the molecule using transmission electron microscopy (TEM). Dilute solutions of 1 in MeCN were placed on a Cu grid, and the solvent carefully evaporated under vacuum. TEM images obtained (Fig. 3a) show uniform nanoparticles with diameters measuring approximately 2.61 nm. These dimensions correspond well with the diameter of the 16-metal ring end of the nano-drum found in the crystal structure. In Fig. 3b a panoramic scanning electron microscopy (SEM) image shows the crystalline nature of 1.
Fig. 1.
Schiff base ligand H2L.
Fig. 2.
Views of the crystal structure of 1 (Eu3+: green; Cd3+: purple).
Fig. 3.
TEM and SEM images of 1.
3.2 Spectroscopic characterization
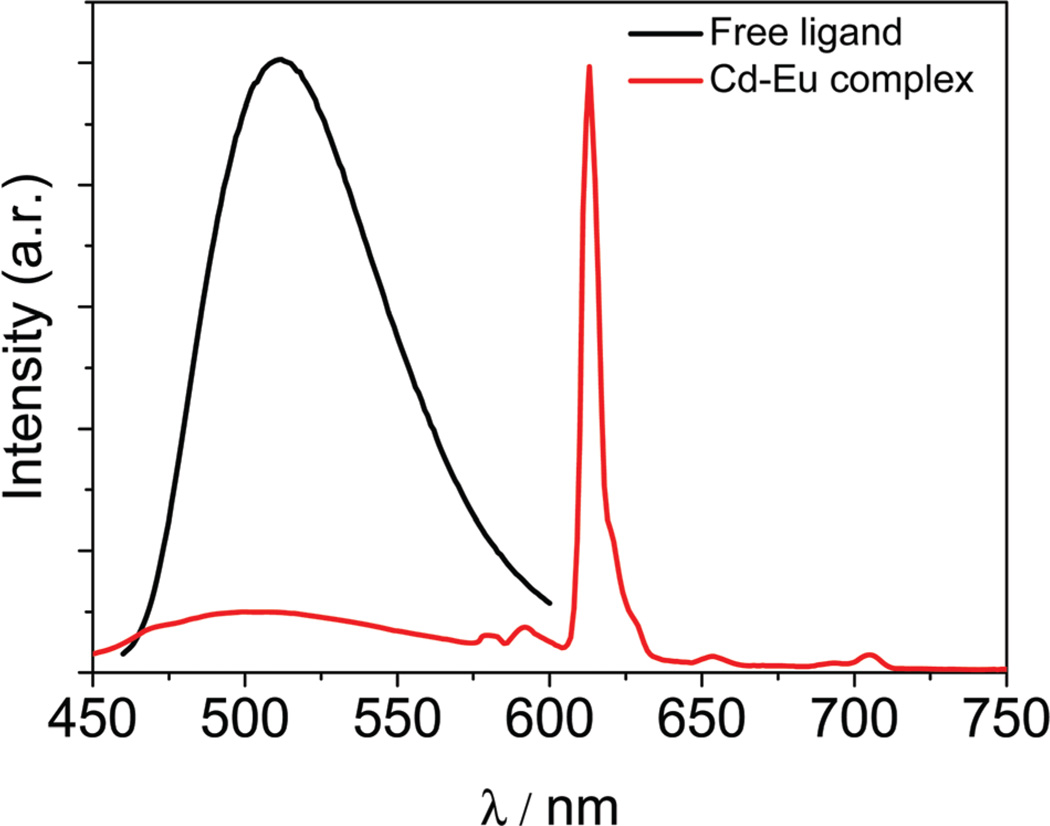
The photophysical properties of the nano-drum 1 were studied in both solution and the solid state. The absorption spectra of the free ligand H2L and 1 in CH3CN are shown in Fig. 4. The free ligand H2L exhibits absorption bands at 226, 260 and 327 nm, which are all red-shifted upon co-ordination to metal ions in 1. The absorption features of 1 are significantly more intense than those of the free ligand (i.e. more than ten times). This is advantageous in terms of sensitizing the lanthanide luminescence. As shown in Fig. 5, for the free ligand H2L, excitation of the absorption band at 295 nm or 410 nm produces a broad emission band at λmax = 515 nm. The excitation spectrum of 1 shows peaks at 320 and 400 nm (Fig. 6). For 1, the typical narrow emission bands of the Eu3+ ion (5D0→7Fj transitions, j = 0, 1, 2, 3 and 4) can be detected upon excitation of the ligand-centered absorption bands in both solution and the solid state at room temperature (Fig. 5). The appearance of the symmetry-forbidden emission 5D0→7F0 at 578 nm is consistent with the Eu3+ ions in the complex occupying sites with low symmetry and having no inversion center, in agreement with the solid state structure.35 A ligand centered (L) 1π–π* emission was detected at 500 nm, a finding consistent with the energy transfer from the ligands to Eu3+ ions being incomplete (100%). This also makes it is difficult to determine accurately the total or intrinsic luminescence quantum yield of 1 due to the mixing of the ligand-centered emission with the lanthanide emission.
Fig. 4.
UV-Vis spectra of free H2L and 1 in CH3CN. (C = 10−7–10−6 M).
Fig. 5.
Emission spectra of the free H2L and 1 in CH3CN. (C = 10−7–10−6 M).
Fig. 6.
Excitation spectrum of 1 in CH3CN. (C = 10−7–10−6 M).
3.3. Cytotoxicity screening
To determine the inherent cytotoxicity of 1 towards potential tumor targets, cell proliferation assays were conducted using an A549 lung cancer cell line and an AGS gastric cancer cell line.
As shown in Fig. 7, nano-drum displayed an IC50 value of 1.37 ± 0.43 µM. A similar IC50 value was obtained for AGS cells, approximately 2 µM. Cd(OAc)2·2H2O and the H2L ligand provided IC50 values of 31.0 ± 7.0 and 29.7 ± 1.2 µM, respectively, while Eu(OAc)3·4H2O displayed no inhibition of cell proliferation under the concentrations tested. On this basis, we conclude that complex 1 may be suitable for use in biological milieus.
Fig. 7.
Dose responsive cell proliferation curves of A549 Lung cells treated with indicated complex. Error bars represent standard deviation.
3.4. Microscopy Studies
Crosslinked polystyrene beads (6 µm) containing 1 (Eu8Cd24L12(OAc)48) were visualized using epifluoresence, TIRF, and two-photon microscopy by observing emission in the visible spectrum (Fig. 8). Evidence that the nano-drum is incorporated intact into the pores of the beads was provided by XPS studies which revealed an Eu : Cd ratio of 1 : 3 consistent with the ratio found in the crystal structure (ESI). Beads loaded separately with 1 and the H2L ligand can both be excited with an illumination source filtered by a DAPI filter and show high signal to background intensity ratio as well as high mean intensity under the DAPI, FITC, TRITC and Cy5 emission filter channels. The use of longer wavelength excitation filters appeared to only slightly excite either sample. Using the TIRF imaging platform, beads loaded with either 1 (Fig. 8c) or H2L (Fig. 8d), exhibited a high fluorescence when excited with a 405 nm laser using identical filter settings and controlled for the same exposure time of 1 frame s−1. The mean intensity of beads loaded with 1 was approximately 67% higher than the H2L loaded beads. Evidence from the solution and solid state photophysical data leads us to suggest that this difference is due to the enhanced fluorescence of 1 compared to H2L. However, the difference in response may also reflect some variations in bead loading concentration and the nature of the layer imaging geometry employed in the TIRF platforms. Improved quantitative measurements of relative fluorescence intensities between bead sets will be necessary to improve the confidence of this observation.
Fig. 8.
Luminex beads doped with 1 and H2L ligand show bright fluorescence when analysed by either epifluorescent or total internal reflection fluorescence (TIRF) microscopy. The nano-drum 1 (A) and H2L ligand (B) can be excited with an illumination source filtered by a DAPI filter. Beads exhibit high fluorescent signals with a 405 nm excitation laser under TIRF imaging for 1 (C) and H2L doped beads (D). All fluorescent images are scaled between a constant minimum and maximum intensity values for comparison. The center wavelengths for the excitation and emission filters are indicated. See methods for details of the filters used.
The two-photon fluorescence images shown in Fig. 9 were acquired at constant laser power and detector sensitivity. Beads were excited at 760 nm, and images shown obtained by PMT with a 535/50 nm emission filter. Beads loaded with H2L ligand were also imaged by two-photon microscopy under similar conditions (ESI Fig. S4†). Control beads lacking 1 did not display significant fluorescence in any of the microscopy studies presented here.
Fig. 9.
Polystyrene beads loaded with 1 imaged by two-photon fluorescence microscopy. (A) Beads were excited at 760 nm, with signal detected in green emission PMT shown at 20× (left panel), 40× (middle panel), and 63× (right panel) magnification. (B) Two-photon excitation spectra for 1 in the visible emission range.
4. Discussion
The presented studies were designed to assess the potential utility of [Eu8Cd24L12(OAc)48] (1), an example of a novel class of molecular lanthanide-based nanoparticles called nano-drums, as a bioprobe for biological imaging applications. Desirable features of a bioprobe include aqueous stability, thermodynamic stability, and favourable photophysical properties, including narrow band emissions and no photobleaching.4,23,36 The crystal structure of 1 confirms that a discrete multinuclear lanthanide nano-drum is formed under the reaction conditions, presumably as the result of a discrete self-assembly process. Spectroscopic characterization of the nano-drum has served to reveal that the physical and chemical properties of complex 1 make it an attractive candidate for the further development into a bioprobe. In the case of 1, the 1H NMR spectra recorded in organic solvents remain unchanged for weeks. This provides support for the notion that they are stable in solution, as well as in crystalline form. In addition, polystyrene beads loaded with the nano-drums of the present study could be stored in aqueous buffer at room temperature for several months, while still retaining their imaging properties, as shown in Fig. 8 and 9.
The photophysical properties of the nano-drum 1 highlight the potential of the complex as an Eu emitter. Moreover, the presence of the Cd/L centre provides added spectral flexibility. Specifically, the centre allows the nano-drum to be used as a non-bleaching fluorescent label that can emit in the visible region of the spectrum and thus be imaged using conventional, commercially available microscopy platforms. Transition metal d-block metal ions introduced into lanthanide complexes may conceivably play two different roles in modulating the luminescence properties of Ln3+ ions. They may enhance the luminescence via d→f energy transfer,32 or they may quench the luminescence via f→d energy transfer.35 Unlike other d metal ions such as Cu2+ and Ni2+, the saturated d10 electronic configuration of the Cd2+ ion prevents quenching of the lanthanide visible luminescence through a d–d transition (f→d energy transfer). With a reasonably high absorption coefficient, the chromogenic Cd(II) components act as sensitizers for lanthanide luminescence in 1.34,36 Interestingly, it was recently reported that there have been no applications of transition metal sensitized lanthanides as cell imaging bioprobes.33 However, were they available such bioprobes would have considerable potential in cell bioimaging applications due to their ability to undergo excitation at wavelengths that would minimize autofluorescence from the intrinsic chromophores present in cells.33 The nano-drum complex 1 is such an example of a chromogenic d-metal component complex that may be photoexcited for imaging. The present report describes efforts aimed at further optimizing the photophysical properties of 1 and carrying out initial cell bioimaging studies.
To determine the inherent cytotoxicity of complexes towards potential tumor targets, cell proliferation assays were conducted using an A549 lung cancer cell line. As shown in Fig. 7, the nano-drum 1 displayed an IC50 value of 1.37 ± 0.43 µM in an A549 lung cancer cell line and 2.0 µM in an AGS gastric cancer cell line. At this point, it is unknown whether the observed cytotoxicity is due to the complex as a whole or due do individual entities within the supramolecular drum assembly (i.e. Cd, Eu, H2L ligand). In this particular cellular assay, the cytotoxicity (antiproliferative activity) of the free metal acetates and ligand (i.e. Eu(OAc)3, Cd(OAc)2 and the H2L ligand) was significantly lower than that of the corresponding nano-drum complex 1. The cadmium salt Cd(OAc)2 and the H2L ligand provided IC50 values of 31.0 ± 7.0 and 29.7 ± 1.2 µM, respectively, while Eu(OAc)3 displayed no inhibition of cell proliferation under the concentrations tested. These observations lead us to suggest that the supramolecular drum assembly is facilitating transport of the ligand and metal components into the cell via an endocytotic mechanism.36 Further studies to substantiate these conclusions are currently underway.
Finally, evaluation of the suitability of the present nano-drums for use in bioimaging applications was assessed using three microscopy platforms configured to detect fluorescence in the visible spectrum. As shown in Fig. 8 and 9, polystyrene beads loaded with 1 could be visualized by epifluorescence, TIRF or two-photon microscopy. Also, intensity scans of the beads (Fig. 8) showed spectral features consistent with previous data obtained in solution. The data also support the notion that excitation can be achieved in the visible region using conventional sources in such a way that minimizes damage to biological materials. Such features are viewed as desirable in situations where ancillary tissue damage might be a genuine concern.
5. Conclusions
We describe here an example of a new class of self-assembling molecular nano-drum (1) that has considerable potential for use in imaging bioprobes and which can be used in range of platforms that rely on fluorescent or luminescent detection. Compared to conventional organic dyes or quantum dots used in biomedical applications, the present 4d–4f molecular cluster does not photobleach or blink. Future studies are focused on creating homogeneous molecular nanoparticles with bioconjugation capabilities. The low temperature, bottom-up, self-assembly route to 1 and related materials should allow for synthetic modification leading to improvements in thermodynamic stability and photophysical properties as well as other important properties, such as lanthanide shielding, solubility, and toxicity.
Supplementary Material
Acknowledgements
This work was supported by the Welch Foundation (F-816) (RAJ), the Ministry of High Education (MOHE), Malaysia under High Impact Research (HIR) – MOHE project (UM.C/625/1/HIR/MoE/CHAN/13/6 H-50001-00-A000034) (to JV and KAB), NIH/NIAID 1U01AI078008-3 and support from the Centre for Blast Injury Study at Imperial College London (to KAB), CPRIT R1003 (to LIRE), NIH-NCI CA68682 and the Welch Foundation (F-1018) (both to JLS), and National Institutes of Health, National Science Foundation, Cancer Prevention Research Institute of Texas, and the Welch Foundation (F1515) (all to EMM). Single crystal X-ray data were collected using instrumentation purchased with funds provided by the National Science Foundation (CHE-0741973). We thank D. Thompson for assistance in editing.
Footnotes
Electronic supplementary information (ESI) available. CCDC 1008697. For ESI and crystallographic data in CIF or other electronic format see DOI: 10.1039/c4dt02646b
References
- 1.Melican K, Richter-Dahlfors A. Biotechnol. J. 2009;4:804. doi: 10.1002/biot.200800347. [DOI] [PubMed] [Google Scholar]
- 2.Tallury P, Malhotra A, Byrne LM, Santra S. Adv. Drug Delivery Rev. 2010;62:424. doi: 10.1016/j.addr.2009.11.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Moore EG, Samuel APS, Raymond KN. Acc. Chem. Res. 2009;42:542. doi: 10.1021/ar800211j. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bünzli J-CG. Chem. Rev. 2010;110:2729. doi: 10.1021/cr900362e. [DOI] [PubMed] [Google Scholar]
- 5.Zhou LJ, Gu ZJ, Liu XX, Yin WY, Tian G, Yan L, Jin S, Ren WL, Xing GM, Li W, Chang XL, Hu ZB, Zhao YL. J. Mater. Chem. 2012;22:966. [Google Scholar]
- 6.Vetrone F, Boyer JC, Capobianco JA, Speghini A, Bettinelli M. Chem. Mater. 2003;15:2737. [Google Scholar]
- 7.Chen XY, Ma E, Liu GK. J. Phys. Chem. 2007;C111:10404. [Google Scholar]
- 8.Shen J, Sun LD, Zhu JD, Wei LH, Sun HF, Yan CH. Adv. Funct. Mater. 2010;20:3708. [Google Scholar]
- 9.Chen DQ, Yu YL, Huang P, Lin H, Shan ZF, Zeng LW, Yang AP, Wang YS. Phys. Chem. Chem. Phys. 2010;12:7775. doi: 10.1039/c003678a. [DOI] [PubMed] [Google Scholar]
- 10.Song YH, Huang YJ, Zhang LH, Zheng YH, Guo N, You HP. RSC Adv. 2012;2:4777. [Google Scholar]
- 11.Zhao Q, Zheng Y, Guo N, Jia Y, Qiao H, Lv W, You HP. CrystEngComm. 2012;14:6659. [Google Scholar]
- 12.Choi S, Yun YJ, Kim SJ, Jung HK. Opt. Lett. 2013;38:1346. doi: 10.1364/OL.38.001346. [DOI] [PubMed] [Google Scholar]
- 13.Song HJ, Yim DK, Roh HS, Cho IS, Kim SJ, Jin YH, Shim HW, Kim DW, Hong KS. J. Mater. Chem. C. 2013;1:500. [Google Scholar]
- 14.Teng X, Zhu YH, Wei W, Wang SC, Huang JF, Naccache R, Hu WB, Tok AIY, Han Y, Zhang QC, Fan QL, Huang W, Capobianco JA, Huang L. J. Am. Chem. Soc. 2012;134:8340. doi: 10.1021/ja3016236. [DOI] [PubMed] [Google Scholar]
- 15.Liu Q, Sun Y, Yang TS, Feng W, Li CG, Li FY. J. Am. Chem. Soc. 2011;133:17122. doi: 10.1021/ja207078s. [DOI] [PubMed] [Google Scholar]
- 16.Mahalingam V, Vetrone F, Naccache R, Speghini A, Capobianco JA. Adv. Mater. 2009;21:4025. [Google Scholar]
- 17.Yi GS, Chow GM. Adv. Funct. Mater. 2006;16:2324. [Google Scholar]
- 18.Chen D, Wang Y. Nanoscale. 2013;5:4621. doi: 10.1039/c3nr00368j. [DOI] [PubMed] [Google Scholar]
- 19.Foucault-Collet A, Gogick KA, White KA, Villette S, Pallier A, Collet G, Kieda C, Li T, Geib SJ, Rosi NL, Petoud S. Proc. Natl. Acad. Sci. U. S. A. 2013;43:17199. doi: 10.1073/pnas.1305910110. PMCID: PMC3808657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Montgomery CP, Murray BS, New EJ, Pal R, Parker D. Acc. Chem. Res. 2009;42:925. doi: 10.1021/ar800174z. [DOI] [PubMed] [Google Scholar]
- 21.Thibon A, Pierre VC. Anal. Bioanal. Chem. 2009;394:107. doi: 10.1007/s00216-009-2683-2. [DOI] [PubMed] [Google Scholar]
- 22.Nishioka T, Fukui K, Matsumoto K. In: Handbook on the Physics and Chemistry of Rare Earths. Gschneidner KA, Bünzli J-CG Jr, Pecharsky VK, editors. Amsterdam: Elsevier Science B. V.; 2007. p. 37. [Google Scholar]
- 23.Eliseeva SV, Bünzli J-CG. Chem. Soc. Rev. 2010;39:189. doi: 10.1039/b905604c. [DOI] [PubMed] [Google Scholar]
- 24.Petoud S, Cohen SM, Bünzli J-CG, Jr, Raymond KN. J. Am. Chem. Soc. 2003;125:13324. doi: 10.1021/ja0379363. [DOI] [PubMed] [Google Scholar]
- 25.Yang X-P, Jones RA, Huang S. Coord. Chem. Rev. 2014;273–274:63–75. [Google Scholar]
- 26.Yang X-P, Schipper D, Jones RA, Holliday BJ, Huang S. J. Am. Chem. Soc. 2013;135:8468. doi: 10.1021/ja4031243. [DOI] [PubMed] [Google Scholar]
- 27.Lam F, Xu J-X, Chan K-S. J. Org. Chem. 1996;61:8414. [Google Scholar]
- 28.Otwinowski Z, Minor W. DENZO-SMN. In: Methods in Enzymology 276: Macromolecular Crystallography. Part A. Carter CWJ, Simon MI, Sweet RM, editors. Academic Press; 1997. p. 307. [Google Scholar]
- 29.Sheldrick GH. SHELX 97, A software package for the solution and refinement of X-ray data. Göttingen, Germany: University of Göttingen; 1997. [Google Scholar]
- 30.Cromer DT, Waber JT. International Tables for X-Ray Crystallography. Vol. 4. Birmingham: Kynoch Press; 1974. table 2.2A. [Google Scholar]
- 31.(a) Meech SR, Philips DJ. J. Photochem. 1983;23:193. [Google Scholar]; (b) Nakamaru K. Bull. Chem Soc. Jpn. 1982;5:2697. [Google Scholar]
- 32.(a) Metcalf DH, Ghirardelli RG, Palmer RA. Inorg. Chem. 1985;24:634. [Google Scholar]; (b) Albin M, Whittle RR, Horrocks WD., Jr Inorg. Chem. 1985;24:4591. [Google Scholar]
- 33.Coogan MP, Fernandez-Moreira V. Chem. Commun. 2014;50:384. doi: 10.1039/c3cc45229h. [DOI] [PubMed] [Google Scholar]
- 34.(a) Ward MD. Coord. Chem. Rev. 2010;254:2634. [Google Scholar]; (b) Vigato RA, Peruzzo V, Tamburini S. Coord. Chem. Rev. 2009;253:1099. [Google Scholar]; (c) Li X-L, Shi L-X, Zhang L-Y, Wen HM, Chen ZN, editors. Inorg. Chem. 2007;46:10892. doi: 10.1021/ic7015676. [DOI] [PubMed] [Google Scholar]; (d) Xu H-B, Shi L-X, Ma E, Zhang L-Y, Wei Q-H, Chen Z-N. Chem. Commun. 2006:1601. doi: 10.1039/b602222g. [DOI] [PubMed] [Google Scholar]; (e) Baca SG, Adams H, Grange CS, Smith AP, Sazanovich I, Ward MD. Inorg. Chem. 2007;46:9779. doi: 10.1021/ic701169c. [DOI] [PubMed] [Google Scholar]; (f) Davies GM, Pope SAJ, Adams H, Faulkner S, Ward MD. Inorg. Chem. 2005;44:4656. doi: 10.1021/ic050512k. [DOI] [PubMed] [Google Scholar]; (g) Yin M, Lei X, Li M, Yuan L, Sun J. J. Phys. Chem. Solids. 2006;67:1372. [Google Scholar]; (h) Xu H-B, Wen H-M, Chen Z-H, Li J, Shi L-X, Chen Z-N. J. Chem. Soc., Dalton Trans. 2010:1948. doi: 10.1039/b919170d. [DOI] [PubMed] [Google Scholar]; (i) Torelli S, Imbert D, Cantuel M, Bernardinelli G, Delahaye S, Hauser A, Bünzli J-CG, Piguet C. Chem. – Eur. J. 2005;11:3228. doi: 10.1002/chem.200401158. [DOI] [PubMed] [Google Scholar]; (j) Piguet C, Bünzli J-CG, Bernardinelli G, Hopfgartner G, Petoud S, Schaad O. J. Am. Chem. Soc. 1996;118:6681. [Google Scholar]; (k) Chi Y-X, Niu S-Y, Jin J, Wang R, Li Y. J. Chem. Soc., Dalton Trans. 2009:7653. doi: 10.1039/b905113a. [DOI] [PubMed] [Google Scholar]; (l) Chi Y-X, Niu S-Y, Wang Z-L, Jin J. Eur. J. Inorg. Chem. 2008:2336–2343. [Google Scholar]
- 35.(a) Wen V, Zhao Y, Wang L, Zhang M, Gao DJ. Rare Earths. 2007;25:679. [Google Scholar]; (b) Brayshaw PA, Bünzli J-CG, Froidevaux P, Harrowfield JM, Kim Y, Sobolev AN. Inorg. Chem. 1995;34:2068. [Google Scholar]; (c) Cantuel M, Bernardinelli G, Imbert D, Bünzli J-CG, Hopfgartner G, Piguet C. J. Chem. Soc., Dalton Trans. 2002:1929. [Google Scholar]; (d) Metcalf DH, Bolender JP, Driver MS, Richardson FS. J. Phys. Chem. 1993;97:553. [Google Scholar]
- 36.(a) Chen Q-Y, Luo Q-H, Hu X-L, Shen M-C, Chen J-T. Chem. – Eur. J. 2002;8:3984. doi: 10.1002/1521-3765(20020902)8:17<3984::AID-CHEM3984>3.0.CO;2-H. [DOI] [PubMed] [Google Scholar]; (b) Wei K-J, Xie Y-S, Ni J, Zhang M, Liu Q-L. Cryst. Growth Des. 2006;6:1341. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.