Figure 2.
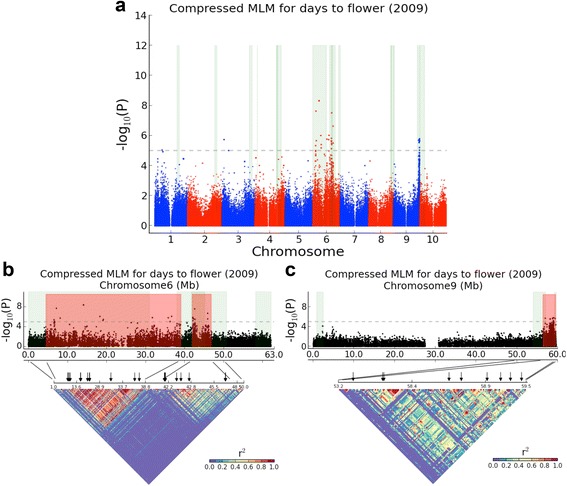
GWAS for flowering time in 2009. (a) Genome-wide Manhattan plot of CMLM. Significance threshold is denoted by the gray dashed line. The 10 sorghum chromosomes are plotted against the negative base-10 logarithm of the association P value. Areas highlighted in green indicate confidence intervals for flowering time determined by QTL mapping. (b) Chromosome Sb06 Manhattan plot of CMLM (top). Red areas show hotspots for 2009 flowering time identified by association mapping. Linkage disequilibrium (r 2 were calculated from SNPs with association p ≤ 0.05 and missing data ≤ 50%) matrices (bottom) are plotted for regions denoted by anchoring lines. Regions of strong LD are shown in red. Significant association markers are denoted by black arrows. (c) Chromosome Sb09 Manhattan plot of CMLM.
