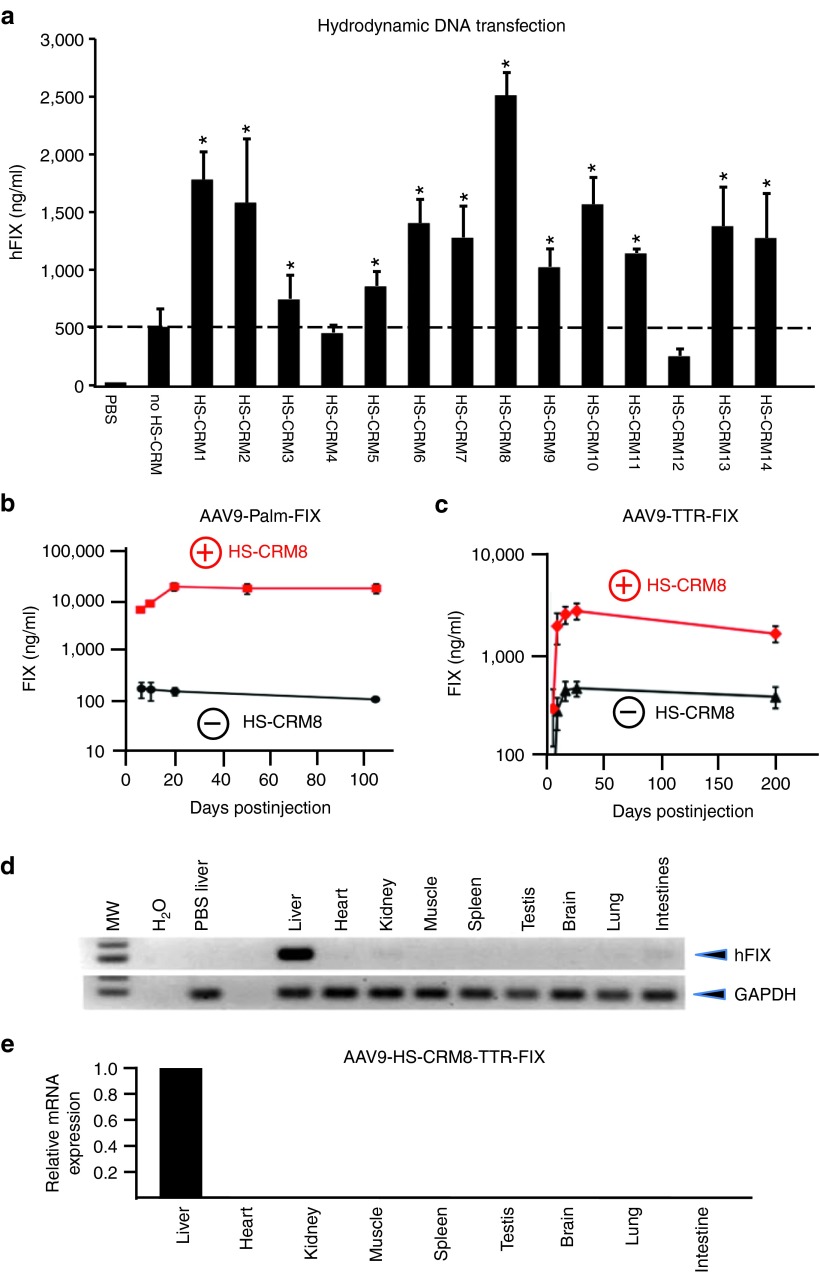
Figure 3.
In vivo validation of HS-CRMs. (a) Semi-high throughput HS-CRM screening in vivo after intravenous hydrodynamic liver-directed injection with pAAV-HS-CRM-TTR-FIX and pAAV-TTR-FIX plasmids at a dose of 2 µg DNA. Significant differences compared to the control without HS-CRM were indicated (t-test, *P ≤ 0.05, mean ± SD). (b) FIX expression levels after intravenous administration of AAV9-HS-CRM8-Palm-FIX and AAV9-Palm-FIX (1 × 1011 vg/mouse) (n = 5 per cohort, C57Bl/6). The difference in FIX expression levels was significant (t-test, *P ≤ 0.0001). (c) FIX expression levels after intravenous administration of AAV9-HS-CRM8-TTR-FIX and AAV9-TTR-FIX control vectors (5 × 109 vg/mouse) (n = 5 per cohort, C57Bl/6). The difference in FIX expression levels was significant (t-test, *P ≤ 0.00005). FIX levels were determined using a hFIX-specific ELISA. (d) Hepatocyte-specificity of AAV9 containing HS-CRM8. RT-PCR analysis on 20 ng total RNA from different organs of C57/Bl6 mice (n = 3) injected intravenously with AAV9-HS-CRM8-TTR-FIX vectors (3 × 1012 vg/mouse); RNA liver samples were amplified with and without RT to exclude genomic DNA amplification. Amplification of hFIX mRNA was not detectable in these control samples without RT (data not shown). (e) Corresponding qRT-PCR analysis of hFIX mRNA levels in the different organs expressed relative to hFIX mRNA levels in the liver. Expression levels (mean ± SD) relative to liver are shown. H2O, water control; MW, molecular weight marker; PBS liver, liver sample of PBS-injected control mice; RT, reverse transcription.