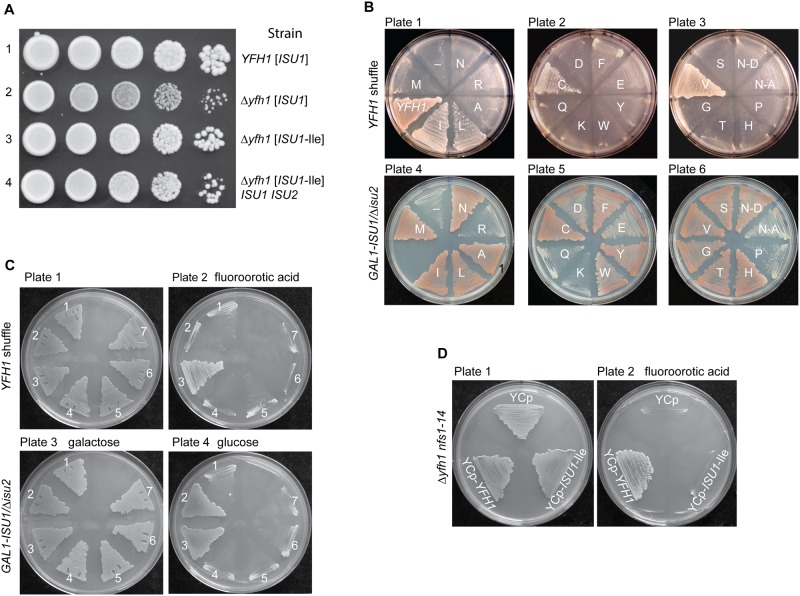
Fig 1. Genetic manipulations of ISU1-Ile and other ISU1 alleles.
(A) Genetic dominance of ISU1-Ile conferring frataxin-bypass. Strains YFH1 [ISU1], Δyfh1 [ISU1], Δyfh1 [ISU1-Ile], and Δyfh1 [ISU1-Ile] ISU1 ISU2 (Table 1) were compared by spotting serial 5-fold dilutions of 105 cells on YPAD plates and photographing three days later. (B) Frataxin-bypass function and scaffold function of ISU1 alleles. A series of plasmids was constructed in the YCplac22 backbone. These plasmids carried native ISU1 or ISU1 alleles with substitutions of residue M141 to all 19 other standard amino acids. N123D or N123A substitutions were also constructed, predicted to give disordered or structured conformations, respectively [34]. Plasmid-borne YFH1 and empty vector (-) were included as controls. To test frataxin-bypass function the ISU1 alleles were transformed into the YFH1 shuffle strain, and the transformed cells were transferred to fluoroorotic acid (FOA) plates to remove the covering YFH1-URA3 plasmid and expose the Δyfh1 phenotype (plates 1–3). To test scaffold function the ISU1 alleles were transformed into the GAL1-ISU1/Δisu2 strain and plated on raffinose carbon source to repress genomic GAL1-ISU1. (C) Effects of cysteine mutations. Plasmids with various cysteine substitutions in Isu1 were tested for frataxin-bypass function by introducing them into the YFH1 shuffle strain and counterselecting on FOA (plates 1 and 2). The Isu1 substitutions were also tested for scaffold function by introducing them into the GAL1-ISU1/Δisu2 strain and shifting the carbon source from galactose to glucose (plates 3 and 4). Key to transformants. 1. Empty plasmid YCplac22, 2. ISU1, 3. ISU1-M141I, 4. ISU1-C69A-M141I, 5. ISU1-C96A-M141I, 6. ISU1-C139A-M141I, 7. ISU1-C139A-M141C. (D) Δyfh1 nfs1-14 double mutant does not support frataxin-bypass by ISU1-M141I. Shuffle strain 113–26 (Δyfh1 nfs1-14 [pRS416-YFH1]) was transformed with YCplac22, YCplac22-YFH1, or YCplac22-ISU1-Ile. Transformants were patched onto tryptophan drop-out medium without (plate 1) or with (plate 2) FOA.