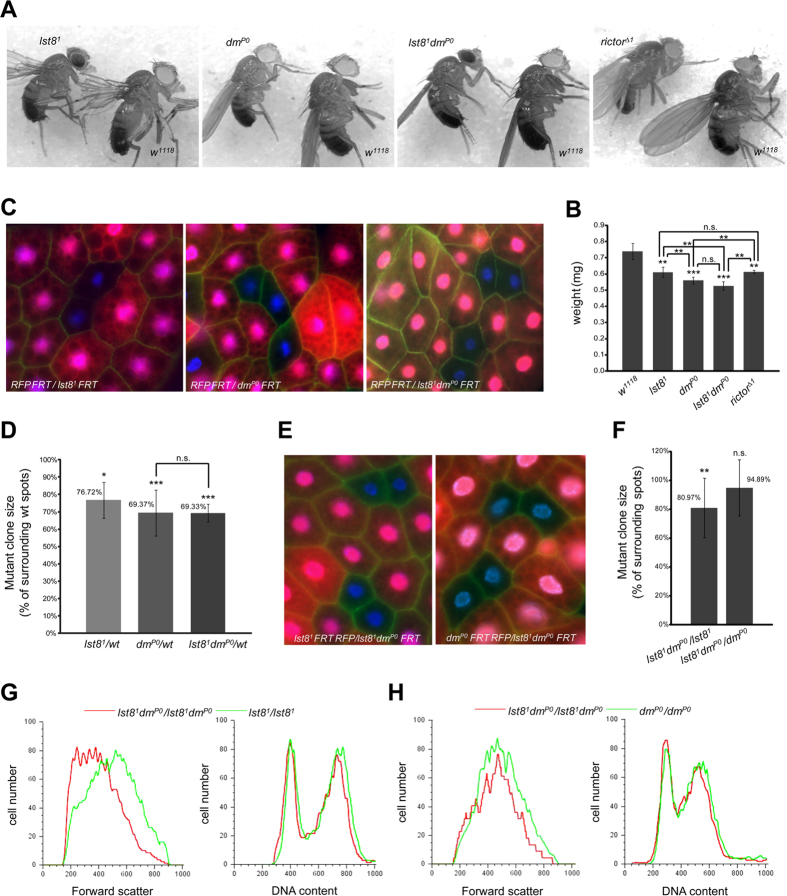
Figure 1. Common pathway for TORC2 and Myc regulated cell growth.
(A) Size comparisons between wild type (w1118) and lst81, dmP0, lst81 dmP0, or rictorΔ1 flies. One-day-old male flies were used for the analysis. (B) Quantification of animal weights. Genotypes of one-day-old male flies are shown. Asterisks indicate statistically significant differences (Student’s unpaired t-test, **p < 0.01; ***p < 0.001; n.s., not significant). (C–F) Comparisons between lst81 dmP0, lst81, and dmP0 fat body cells. (C) The FLP/FRT system was used to generate mosaic lst81, dmP0, or lst81 dmP0 mutant clones marked by the absence of RFP (hs-flp ub-RFP FRT/lst81 FRT, hs-flp ub-RFP FRT/dmP0 FRT, and hs-flp ub-RFP FRT/lst81 dmP0 FRT animals, respectively) (red, RFP; green, phalloidin; blue, DAPI). (D) Relative size of lst81, dmP0, or lst81 dmP0 cells compared with controls. Asterisks indicate statistically significant differences from wild-type cells (Student’s unpaired t-test, ***p < 0.001; n.s., not significant). (E) The FLP/FRT system was used to generate mosaic lst81 dmP0 double mutant clones in larval fat bodies of lst81 hs-flp ub-RFP FRT/lst81 dmP0 FRT and dmP0 hs-flp ub-RFP FRT/lst81 dmP0 FRT animals, respectively. The lst81 dmP0 mutant cells are marked by the absence of RFP (red, RFP; green, Phalloidin; blue, DAPI). (F) Relative sizes of lst81 dmP0 double mutant cells compared with lst81 or dmP0 cells. Asterisks indicate statistically significant differences from single mutant cells (Student’s unpaired t-test, **p < 0.01; n.s., not significant). (G and H) Flow cytometry was performed on dissociated wing discs from lst81 hs-flp ub-RFP FRT/lst81 dmP0 FRT and dmP0 hs-flp ub-RFP FRT/lst81 dmP0 FRT animals. Cells lacking lst8 (lst81/lst81; green trace in G), dm (dmP0/dmP0; green trace in H), or lst8 and dm (lst81 dmP0/lst81 dmP0; red trace) were compared. Hoechst 33342 staining was used to assess DNA content (right), and FSC was used to quantify cell size (left).