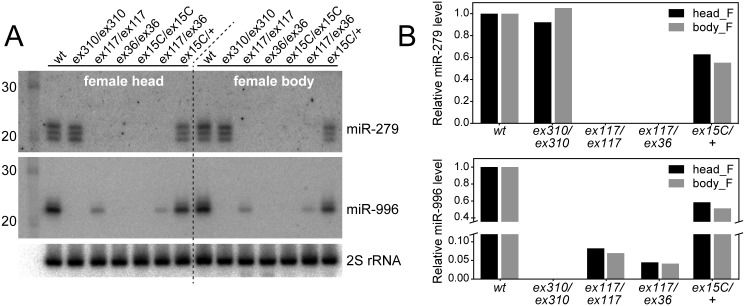
Fig 2. Severe loss of mature miR-996 expression in mir-279 deletion alleles.
(A) Northern blots of miR-279 and miR-996 in various mir-279 and mir-996 homozygous or trans-heterozygous allele combinations. In mir-279 alleles [ex117] and [ex36] that retain the mir-996 genomic DNA, the levels of mature miR-996 are strongly diminished ([ex117]) or nearly undetectable ([ex36]). mir-996[ex310] is a deletion of the mir-996 region that does not affect mir-279 and mir-279/996[ex15C] deletes both miRNAs. (B) Quantifications of mature miR-279 and miR-996 levels. Homozygous [ex117] mutants expressed ~10% of the wild type level of miR-996 and [ex117/ex36] transheterozygous mutants expressed ~5% of miR-996. Note that the expression level for both miRNAs is copy-number dependent, since heterozygous mir-279/996[ex15C] flies expressed roughly half of both miRNAs compared to wild type animals.