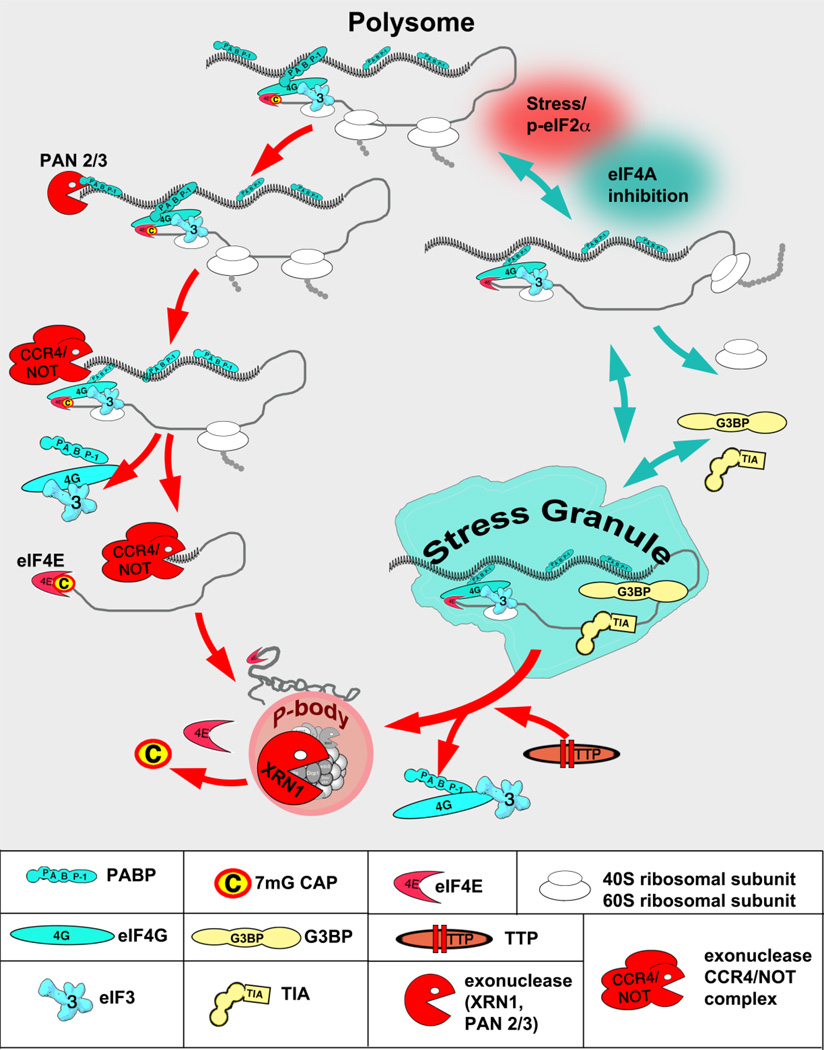
Figure 1. RBP-RNA granules interplay on the target mRNA.
Under steady-state conditions, mRNA is actively engaged into translation as a part of polysome and in the form of translationally-competent mRNP consisting of several translation initiation factors (such as cap-binding complex eIF4F and multi-subunit eIF3) and mRNA-bound RBPs (such as poly(A)-bound PABP). In response to cell-intrinsic and extracellular changes mRNA translation is arrested and transcript can be routed from the polysomes to SGs (blue arrows) or PBs (red arrows). Transcripts destined for assembly into PBs are first deadenylated by one or more mRNA deadenylases (PAN2/3 and CCR4/NOT are shown as examples). This causes the release of PABP molecules from the mRNA poly(A) tail, and the remodeling/decircularization of the mRNP which primes it for decapping. PB-associated decay enzymes subsequently may mediate mRNA decapping and decay within PBs. In contrast, transcripts assembled into SGs retain their poly(A) tails and associated PABP. Stresses, which may trigger phosphorylation of eIF2α (red shading) or not (such as inhibition of eIF4A, aqua shading), disrupt translational initiation of the mRNA, resulting in the runoff of ribosomes and the accumulation of stalled mRNPs. Binding of mRNAs by aggregation-prone RNA binding proteins (such as TIA-1, G3BP1) subsequently promotes its coalesce into SGs. Certain decay-promoting proteins (TTP, BRF1) can promote deadenylation and decay of SG-associated transcripts in tandem with promoting SG-PB fusion/interactions. Multiple additional proteins and signaling molecules associate with SGs or PBs once they are formed, depending on conditions.