Table 1.
Differentially abundant proteins during E. arvense spore germination.
| Spot No.(a) | Protein name(b) | Subcellular location(c) | Plant species(d) | Gi No.(e) | Thr. MW(Da)/pI(f) | Exp. MW(Da)/pI(g) | Cov (%)(h) | Sco(i) | QM(j) | V% ± SD(k) MS RS DCS GS SPC |
|---|---|---|---|---|---|---|---|---|---|---|
| PHOTOSYNTHESIS (17) | ||||||||||
| 121 | Chlorophyll a/b binding protein (CAB) | Chl | Hedera helix | 12,582 | 20,759/4.83 | 25,376/4.71 | 10 | 57 | 3 | 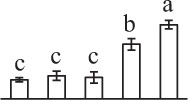 |
| 346 | Ribulose-1,5-bisphosphate carboxylase/oxygenase large subunit (RBCL) | Chl | Eryngium bourgatii | 1,292,976 | 53,093/5.56 | 28,686/6.28 | 6 | 59 | 3 | 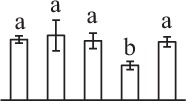 |
| 763 | Ribulose-1,5-bisphosphate carboxylase/oxygenase large subunit (RBCL) | Chl | Donatia fascicularis | 1,304,292 | 49,896/6.32 | 134,736/5.48 | 8 | 55 | 5 | 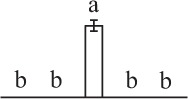 |
| 902 | Ribulose-1,5-bisphosphate carboxylase/oxygenase large subunit (RBCL) | Chl | Grammitis diminuta | 340,031,166 | 45,938/6.26 | 50,502/6.31 | 5 | 61 | 2 | 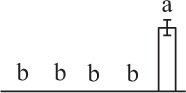 |
| 117 | Ribulose-1,5-bisphosphate carboxylase/oxygenase large subunit (RBCL) | Chl | Equisetum telmateia | 16,565,336 | 48,750/6.26 | 52,974/6.47 | 15 | 57 | 7 | 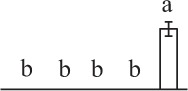 |
| 196 | Ribulose-1,5-bisphosphate carboxylase/oxygenase large subunit (RBCL) | Chl | E. telmateia | 16,565,336 | 48,750/6.26 | 51,502/6.47 | 16 | 52 | 6 | 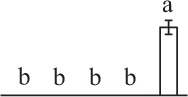 |
| 194 | Ribulose-1,5-bisphosphate carboxylase/oxygenase large subunit (RBCL) | Chl | E. telmateia | 16,565,336 | 48,750/6.26 | 51,418/6.33 | 13 | 53 | 6 | 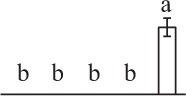 |
| 400 | Ribulose-1,5-bisphosphate carboxylase/oxygenase large subunit (RBCL) | Chl | E. telmateia | 16,565,336 | 48,750/6.26 | 49,452/6.31 | 11 | 53 | 5 | 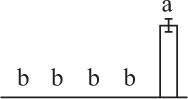 |
| 23 | Ribulose-1,5-bisphosphate carboxylase/oxygenase large subunit (RBCL) | Chl | E. bourgatii | 1,292,976 | 53,093/5.56 | 50,872/5.02 | 4 | 78 | 2 | 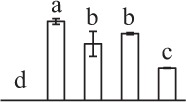 |
| 528 | Ribulose-1,5-bisphosphate carboxylase/oxygenase large subunit (RBCL) | Chl | Equisetum arvense | 1,352,773 | 52,493/5.86 | 67,620/4.64 | 7 | 60 | 3 | 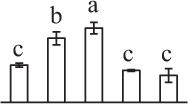 |
| 100 | Ribulose-1,5-bisphosphate carboxylase/oxygenase large subunit (RBCL) | Chl | Isoetes capensis | 83,032,384 | 47,561/6.30 | 42,425/5.47 | 6 | 55 | 3 | 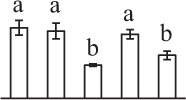 |
| 496 | Ribulose-1,5-bisphosphate carboxylase/oxygenase large subunit-binding protein subunit beta (RBP) | Chl | Solanum lycopersicum | 460,379,814 | 64,527/5.46 | 71,249/5.21 | 4 | 61 | 2 | 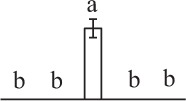 |
| 34 | Ribulose-1,5-bisphosphate carboxylase/oxygenase activase (RCA) | Chl | Gossypium hirsutum | 12,620,883 | 48,609/5.06 | 48,999/5.19 | 11 | 98 | 4 | 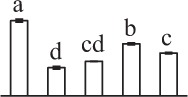 |
| 184 | Ribulose-1,5-bisphosphate carboxylase/oxygenase activase (RCA) | Chl | Hordeum vulgare | 100,614 | 47,496/5.64 | 47,356/6.31 | 11 | 94 | 4 | 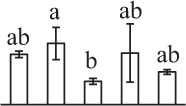 |
| 125 | Ribulose-1,5-bisphosphate carboxylase/oxygenase activase (RCA) | Chl | Musa acuminata subsp. malaccensis | 695,062,479 | 47,898/6.18 | 42,859/5.67 | 8 | 75 | 3 | 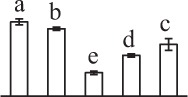 |
| 308 | Transketolase (TK) | Chl | Spinacia oleracea | 2,529,342 | 80,744/6.20 | 95,817/6.20 | 8 | 156 | 5 | 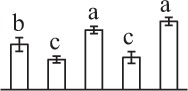 |
| 84 | Transketolase (TK) | Chl | S. lycopersicum | 460,388,792 | 80,615/6.26 | 85,056/6.11 | 8 | 90 | 7 | 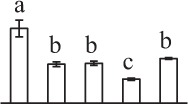 |
| CARBOHYDRATE AND ENERGY METABOLISM (9) | ||||||||||
| 685 | Unknown, pyruvate dehydrogenase E1 component subunit beta* (PDH) | Chl | Glycine max | 255,647,166 | 44,596/6.28 | 27,410/6.46 | 4 | 60 | 2 | 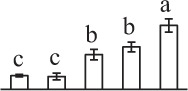 |
| 160 | Malate dehydrogenase (MDH) | #Chl, Cyt, Mit, Pox | Arabidopsis thaliana | 15,219,721 | 35,890/6.11 | 46,411/4.98 | 12 | 74 | 3 | 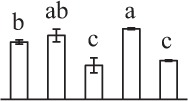 |
| 503 | Malate dehydrogenase (MDH) | #Chl, Cyt, Mit, Pox | A. thaliana | 15,219,721 | 35,890/6.11 | 42,425/5.53 | 6 | 57 | 2 | 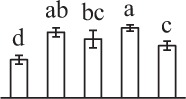 |
| 260 | Malate dehydrogenase (MDH) | #Chl, Cyt, Mit, Pox | A. thaliana | 11,133,509 | 35,548/6.11 | 29,729/6.41 | 6 | 59 | 3 | 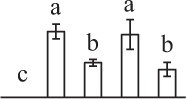 |
| 164 | Malate dehydrogenase (MDH) | #Chl, Cyt, Mit, Pox | Beta vulgaris subsp. vulgaris | 731,361,010 | 41,677/5.74 | 36,353/5.84 | 12 | 193 | 4 | 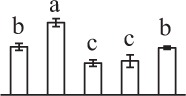 |
| 163 | Malate dehydrogenase (MDH) | Chl | Brachypodium distachyon | 357,147,942 | 41,864/6.97 | 38,560/5.94 | 10 | 60 | 4 | 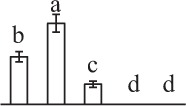 |
| 486 | Enolase | Cyt | Tarenaya hassleriana | 729,317,446 | 51,639/5.91 | 48,823/5.62 | 3 | 54 | 2 | 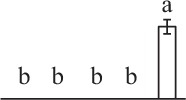 |
| 489 | 6-phosphogluconate dehydrogenase (6-PGDH) | #Chl, Cyt | G. max | 356,513,305 | 54,116/6.25 | 24,817/5.21 | 12 | 53 | 5 | 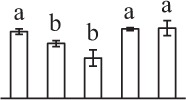 |
| 192 | Fructokinase (FK) | Chl | Lycopersicon esculentum | 23,476,263 | 40,620/5.41 | 37,710/4.91 | 8 | 129 | 3 | 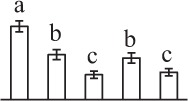 |
| OTHER METABOLISMS (5) | ||||||||||
| 135 | Enoyl-acyl carrier protein reductase (EAR) | Chl | Oryza sativa subsp. japonica | 75,225,229 | 39,277/8.81 | 32,651/6.16 | 5 | 21 | 2 |  |
| 197 | Hypothetical protein PHAVU_001G035500g, glycine decarboxylase* (GDC) | Mit | Phaseolus vulgaris | 593,795,946 | 116,167/6.65 | 117,088/6.30 | 11 | 157 | 12 | 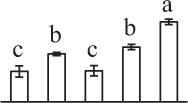 |
| 362 | 3-isopropylmalate dehydrogenase (IPMDH) | Chl | A. thaliana | 15,241,338 | 44,305/5.75 | 28,512/4.87 | 23 | 67 | 7 |  |
| 445 | Hypothetical protein SELMODRAFT_406755, containing cd00517 ATP-sulfurylase domain* (ATPS) | Chl | Selaginella moellendorffii | 302,763,978 | 56,179/6.69 | 26,370/5.59 | 5 | 60 | 3 | 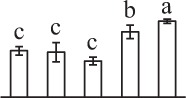 |
| 190 | Predicted protein, pyridoxal biosynthesis protein PDX1* | Cyt | Physcomitrella patens | 168,019,502 | 33,769/6.03 | 29,893/5.82 | 15 | 324 | 8 | 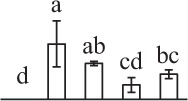 |
| SIGNALING AND VESICLE TRAFFICKING (8) | ||||||||||
| 282 | Unknown, containing PLN02804 chalcone isomerase domain* (CHI) | Chl | Picea sitchensis | 116,784,316 | 23,688/5.23 | 23,141/4.75 | 7 | 56 | 2 | 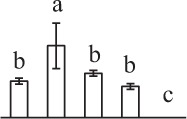 |
| 1063 | Hypothetical protein SELMODRAFT_151778, containing cd00200 WD40 domain* (WD40) | Nuc | S. moellendorffii | 302,791,020 | 37,267/5.65 | 36,189/5.74 | 6 | 84 | 2 | 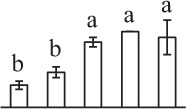 |
| 207 | 14-3-3 protein | Cyt | S. oleracea | 440,573,600 | 30,029/4.84 | 27,277/4.79 | 16 | 79 | 6 | 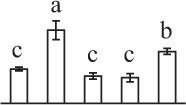 |
| 276 | Unknown, containing cd00877 Ran GTPase domain* (RAN) | Nuc | P. sitchensis | 116,794,384 | 25,374/6.30 | 27,727/6.65 | 20 | 52 | 4 | 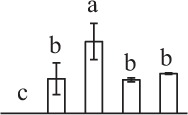 |
| 317 | GTP-binding nuclear protein Ran/TC4 (RAN) | Nuc | Vicia faba | 585,783 | 25,274/6.39 | 27,750/6.73 | 27 | 187 | 6 | 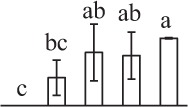 |
| 340 | Unknown, containing cd00877 Ran GTPase domain* (RAN) | Nuc | P. sitchensis | 116,794,384 | 25,374/6.30 | 30,115/5.59 | 24 | 53 | 5 | 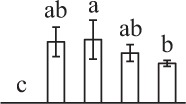 |
| 764 | Unknown, containing cd00877 Ran GTPase domain* (RAN) | Nuc | P. sitchensis | 116,794,384 | 25,374/6.30 | 44,900/6.58 | 30 | 51 | 6 | 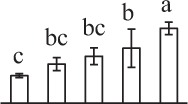 |
| 123 | ADP-ribosylation factor (ARF) | Gol | Chlamydomonas reinhardtii | 1,703,374 | 20,747/6.92 | 19,121/6.71 | 29 | 92 | 4 | 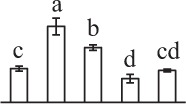 |
| CELL STRUCTURE (5) | ||||||||||
| 397 | Os03g0718100, actin* | Cyt | O. sativa subsp. japonica | 115,454,971 | 42,014/5.30 | 32,585/4.95 | 9 | 82 | 3 | 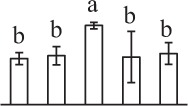 |
| 554 | Reversibly glycosylated polypeptide (RGP) | Cyt | Ricinus communis | 223,546,230 | 41,557/5.82 | 39,973/5.41 | 14 | 105 | 5 | 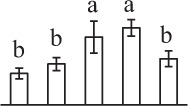 |
| 118 | Reversibly glycosylated polypeptide (RGP) | Cyt | Solanum tuberosum | 34,582,499 | 42,146/5.71 | 35,827/5.36 | 24 | 74 | 8 | 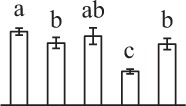 |
| 299 | Unknown protein, rhamnose biosynthetic enzyme 1* (RBE) | #Cyt | A. thaliana | 8,493,590 | 33,861/5.73 | 29,119/6.48 | 7 | 127 | 4 | 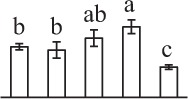 |
| 482 | Caffeoyl-CoA O-methyltransferase (CCoAOMT) | #Cyt | Eucalyptus gunnii | 3,023,419 | 28,010/5.02 | 29,0934.81 | 11 | 73 | 2 | 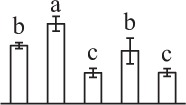 |
| CELL CYCLE (2) | ||||||||||
| 104 | Cell division cycle protein 48 homolog (CDC48) | Cyt | T. hassleriana | 729,396,339 | 89,888/5.09 | 103,252/5.22 | 21 | 119 | 14 | 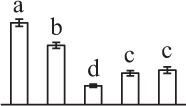 |
| 139 | Proliferation-associated protein 2G4-like (PA2G4) | Nuc | Nicotiana tomentosiformis | 697,180,533 | 43,837/5.96 | 55,758/6.73 | 5 | 52 | 2 | 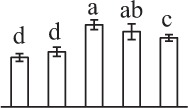 |
| TRANSCRIPTION RELATED (1) | ||||||||||
| 938 | Predicted protein, containing cd00771 threonyl-tRNA synthetase class II core catalytic domain* (ThrRS) | Cyt | P. patens | 168,012,416 | 71,002/5.96 | 80,637/6.37 | 3 | 54 | 2 | 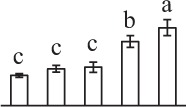 |
| PROTEIN SYNTHESIS (10) | ||||||||||
| 83 | Predicted protein, 40S ribosomal protein SA (RPSA) | Cyt | P. patens | 168,017,628 | 31,539/4.72 | 43,804/4.58 | 7 | 101 | 2 | 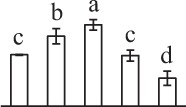 |
| 454 | Hypothetical protein AMTR_s00087p00135370, containing cd08065 eukaryotic translation initiation factor 3* (eIF3) | #Chl, Cyt, Mit, Nuc | Amborella trichopoda | 548,842,775 | 44,710/5.17 | 39,587/4.64 | 5 | 82 | 2 | 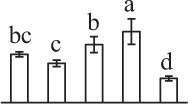 |
| 66 | Hypothetical protein, eukaryotic initiation factor 4A* (eIF4A) | Cyt | P. patens | 168,026,095 | 47,119/5.46 | 48,999/5.32 | 25 | 162 | 11 | 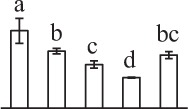 |
| 878 | Eukaryotic initiation factor 4A (eIF4A) | Cyt | Nicotiana. tabacum | 1,170,511 | 47,098/5.37 | 49,134/5.23 | 14 | 118 | 6 | 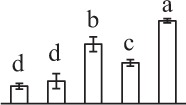 |
| 94 | OSJNBa0020P07.3, elongation factor 2* (EF2) | Cyt | O. sativa subsp. japonica | 38,344,860 | 94,939/5.85 | 115,425/6.42 | 9 | 111 | 7 | 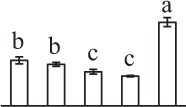 |
| 538 | Hypothetical protein SELMODRAFT_411087, elongation factor 2* (EF2) | Cyt | S. moellendorffii | 302,773,640 | 94,568/6.00 | 114,195/6.33 | 5 | 167 | 5 | 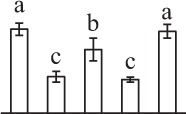 |
| 510 | Hypothetical protein SELMODRAFT_143627, elongation factor G* (EF-G) | #Chl, Mit | S. moellendorffii | 302,765,284 | 75,483/5.31 | 119,422/6.41 | 6 | 75 | 4 | 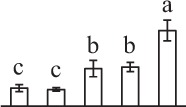 |
| 942 | Elongation factor Tu (EF-Tu) | Mit | Setaria italica | 514,812,465 | 48,530/5.99 | 44,724/6.18 | 16 | 63 | 8 | 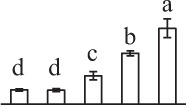 |
| 205 | Uncharacterized protein LOC105056625, containing cd14275 elongation factor Ts domain* (EF-Ts) | Chl | Elaeis guineensis | 743,840,139 | 125,269/4.92 | 27,102/5.82 | 2 | 130 | 5 |  |
| 625 | Tyrosine phosphorylated protein A (TypA) | Chl | Suaeda salsa | 162,424,768 | 75,474/6.71 | 85,342/5.71 | 6 | 205 | 4 | 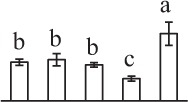 |
| PROTEIN FOLDING AND PROCESSING (8) | ||||||||||
| 542 | Hypothetical protein SELMODRAFT_440382, containing pfam00012 heat shock protein 70 domain* (HSP70) | Cyt | S. moellendorffii | 302,770,212 | 71,931/5.17 | 80,237/5.18 | 13 | 295 | 7 | 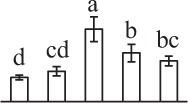 |
| 1002 | Heat shock protein 70 (HSP70) | Cyt | Populus trichocarpa | 224,098,390 | 71,620/5.14 | 80,105/5.21 | 16 | 135 | 8 | 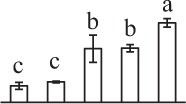 |
| 1566 | Heat shock protein 70 (HSP70) | Cyt | Petunia × hybrida | 20,559 | 71,137/5.07 | 106,146/6.34 | 4 | 92 | 2 | 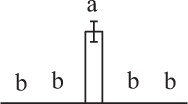 |
| 314 | Heat shock protein 70 (HSP70) | Cyt | Dactylis glomerata | 188,011,548 | 72,002/5.03 | 86,310/5.38 | 21 | 338 | 13 | 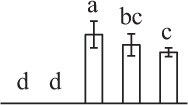 |
| 928 | Heat shock protein 70 like protein (HSP70) | Cyt | S. lycopersicum | 460,394,037 | 72,308/5.16 | 91,122/5.25 | 7 | 122 | 5 | 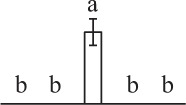 |
| 598 | Predicted protein, containing pfam00183 heat shock protein 90 domain* (HSP90) | Cyt | P. patens | 168,034,606 | 79,652/4.93 | 87,379/5.25 | 5 | 89 | 3 | 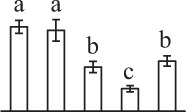 |
| 259 | T-complex protein 1 subunit alpha (TCP1α) | Cyt | A. thaliana | 135,535 | 59,477/5.93 | 66,881/6.22 | 8 | 246 | 4 | 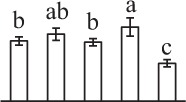 |
| 106 | Hypothetical protein VITISV_000290, T-complex protein 1 subunit gamma* (TCP1γ) | Cyt | V. vinifera | 147,784,740 | 61,064/6.06 | 72,989/6.14 | 5 | 94 | 3 | 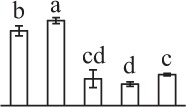 |
| PROTEIN DEGRADATION (8) | ||||||||||
| 284 | Alpha7 proteasome subunit (PSA7) | Cyt, Nuc | N. tabacum | 14,594,925 | 27,466/6.11 | 25,449/5.20 | 8 | 79 | 2 | 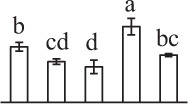 |
| 199 | Zinc dependent protease (ZDP) | Chl | Trifolium pratense | 84,468,286 | 74,746/5.82 | 73,106/5.66 | 12 | 106 | 7 | 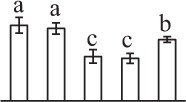 |
| 227 | Zinc dependent protease (ZDP) | Chl | T. pratense | 84,468,286 | 74,746/5.82 | 26,150/5.94 | 10 | 87 | 7 | 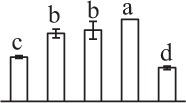 |
| 37 | Zinc dependent protease (ZDP) | Chl | T. pratense | 84,468,286 | 74,746/5.82 | 72,520/5.57 | 5 | 93 | 3 | 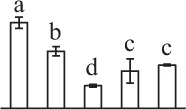 |
| 869 | Zinc metalloprotease (ZMP) | Chl, Mit | A. thaliana | 10,120,424 | 121,539/5.39 | 131,816/5.10 | 1 | 44 | 2 | 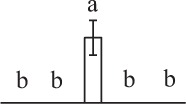 |
| 354 | Predicted protein, containing pfam06480 FtsH extracellular domain* (FtsH) | Chl | Micromonas pusilla CCMP1545 | 303,275,720 | 77,421/5.29 | 75,882/5.26 | 6 | 113 | 4 | 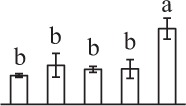 |
| 508 | Predicted protein, ATP-dependent zinc metalloprotease FtsH* (FtsH) | Chl | P. patens | 168,001,910 | 68,933/5.23 | 108,174/5.50 | 3 | 75 | 2 | 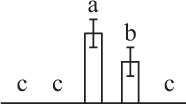 |
| 59 | ATP-dependent Clp protease ATP-binding subunit ClpC (CLPC) | Chl | A. thaliana | 9,758,239 | 103,616/6.36/ | 94,506/5.45/ | 9 | 415 | 9 | 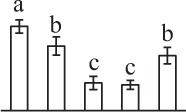 |
| STRESS AND DEFENSE (7) | ||||||||||
| 414 | Hypothetical protein MIMGU_mgv1a021611mg, containing cd03015 2-cys peroxiredoxin domain* (Prx) | Chl, Cyt | Erythranthe guttata | 604,334,612 | 21,153/4.98 | 21,812/4.68 | 17 | 132 | 4 | 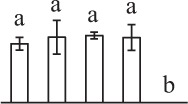 |
| 92 | 2-cys peroxiredoxin-like protein (Prx) | Chl, Cyt | Hyacinthus orientalis | 47,027,073 | 21,956/4.93 | 22,930/5.24 | 12 | 58 | 3 | 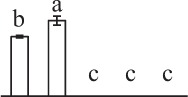 |
| 166 | Ascorbate peroxidase (APX) | Cyt | Eucalyptus grandis | 702,241,628 | 27,613/6.07 | 25,671/5.66 | 7 | 104 | 2 | 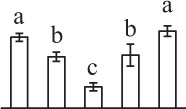 |
| 244 | Dehydroascorbate reductase-like protein (DHAR) | Cyt | S. tuberosum | 76,573,291 | 23,610/6.32 | 24,534/5.60 | 11 | 147 | 3 | 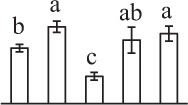 |
| 361 | Dehydroascorbate reductase-like protein (DHAR) | Cyt | S. tuberosum | 76,573,291 | 23,610/6.32 | 24,783/5.64 | 11 | 105 | 3 | 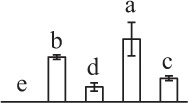 |
| 899 | Dehydroascorbate reductase-like protein (DHAR) | Cyt | S. tuberosum | 76,160,951 | 23,596/6.09 | 44,783/6.60 | 11 | 92 | 3 | 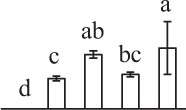 |
| 294 | Chloroplast drought-induced stress protein of 32 kDa (CDSP32) | Chl, Cyt | S. tuberosum | 2,582,822 | 33,779/8.07 | 31,436/6.26 | 8 | 87 | 3 | 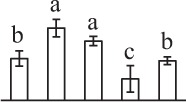 |
Assigned spot number as indicated in Figure 3.
The name and functional category of the proteins identified by ESI-Q-TOF and ESI-Q-Trap tandem mass spectrometry. Protein names marked with an asterisk (*) have been edited by us depending on searching against NCBI non-redundant protein database for functional domain. The abbreviations for the protein names are indicated in the bracket after protein names.
Protein subcellular localization predicted by softwares (YLoc, LocTree3, Plant-mPLoc, ngLOC, and TargetP). Only the consistent predictions from at least two tools were accepted as a confident result. Pounds (#) indicate prediction results were inconsistent among five tools. The subcellular localizations were predicted based on literature listed in Supplementary Table S4. Chl, chloroplast; Cyt, cytoplasm; Gol, Golgi apparatus; Mit, mitochondria; Nuc, nucleus; Pox, peroxisome.
The plant species that the peptides matched from.
Database accession number from NCBI non-redundant protein database.
Theoretical (f) and experimental (g) molecular weight (Da) and pI of identified proteins. Theoretical values were retrieved from the protein database. Experimental values were calculated using ImageMaster 2D version 5.0.
The amino acid sequence coverage for the identified proteins.
The Mascot score obtained after searching against the NCBI non-redundant protein database.
The number of matched peptides for each protein.
The mean values of protein spot volumes relative to total volume of all the spots. Five spore germination stages, MS, mature spores; RS, rehydrated spores, DCS, double-celled spores, GS, germinated spores; and SPC, spores with protonemal cells were performed. Error bar indicates ± standard deviation (SD). Letters indicate statistically significant differences (p <0.05) among five stages of spore germination as determined by One-Way ANOVA.
