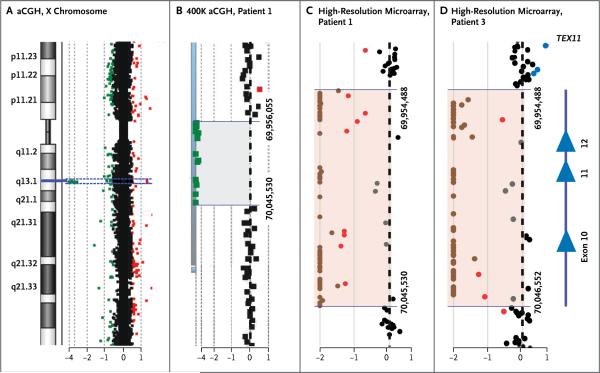
Figure 1. Hemizygous Deletion of TEX11 Exons 10 to 12 and Flanking Intronic Regions in Two Men with Azoospermia.
Panel A shows the array comparative genomic hybridization (aCGH) plot of the X chromosome. On the left, an idiogram of the X chromosome shows a region of interest (blue line) at the Xq13.2 band. On the right, dots represent oligonucleotide DNA probes, arranged according to their physical map locations from the distal p arm (top) to the distal q arm (bottom) of the X chromosome. For each probe, the fluorescence intensity of the test signal relative to the reference signal is converted to a logarithmic (log2) value and shown along the x axis. Probes with a log2 ratio clustered around zero (black dots) indicate DNA segments with a normal copy number. A positive log2 ratio (above +0.3 [red dots]) indicates a gain (extra copy) of the chromosomal region, whereas intervals with a negative log2 ratio (below −0.5 [green dots]) represent a loss (deletion) in DNA copy number. Panel B shows a magnified view of the deleted region detected by 12 probes with the use of the 400K whole-genome array (gray shaded area). Genomic coordinates (GRCh37/hg19 assembly) of the deletion start at 69,956,055 and stop at 70,045,530 nucleotides. Panels C and D show that the aCGH plot of the 180K oligonucleotide X-chromosome high-resolution microarray more precisely delineates the deletion intervals in Patients 1 and 3. Chromosomal alterations suggestive of a homozygous or heterozygous deletion are indicated by brown and red dots, respectively. In a sample obtained from Patient 1 (Panel C), the deletion spans chrX:69,954,488 to 70,045,530. In a sample obtained from Patient 3 (Panel D), the deletion spans chrX:69,954,488 to 70,046,552 regions (pink shaded areas). On the right side of Panel D, Xq13.2 deletions in both patients encompass exons 10 to 12 (blue triangles) of TEX11.