Abstract
The nuclear interferon-inducible-16 (IFI16) protein acts as DNA sensor in inflammasome signaling and as viral restriction factor. Following Herpesvirus infection or UV-B treatment, IFI16 delocalizes from the nucleus to the cytoplasm and is eventually released into the extracellular milieu. Recently, our group has demonstrated the occurrence of IFI16 in sera of systemic-autoimmune patients that hampers biological activity of endothelia through high-affinity membrane binding. As a continuation, we studied the activity of endotoxin-free recombinant IFI16 (rIFI16) protein on primary endothelial cells. rIFI16 caused dose/time-dependent upregulation of IL-6, IL-8, CCL2, CCL5, CCL20, ICAM1, VCAM1, and TLR4, while secretion of IL-6 and IL-8 was amplified with lipopolysaccharide synergy. Overall, cytokine secretion was completely inhibited in MyD88-silenced cells and partially by TLR4-neutralizing antibodies. By screening downstream signaling pathways, we found that IFI16 activates p38, p44/42 MAP kinases, and NF-kB. In particular, activation of p38 is an early event required for subsequent p44/42 MAP kinases activity and cytokine induction indicating a key role of this kinase in IFI16 signaling. Altogether, our data conclude that extracellular IFI16 protein alone or by synergy with lipopolysaccharide acts like Damage-associated molecular patterns propagating “Danger Signal” through MyD88-dependent TLR-pathway.
Introduction
“IFN signature” is a very common feature of different autoimmune diseases (Bennett and others 2003; Sozzani and others 2010; Higgs and others 2011; Ronnblom and others 2011; Ruiz-Riol and others 2011; Bronson and others 2012; Forster 2012; Ronnblom and Eloranta 2013), as these cytokines have been identified as major contributors in the initiation and progression of an autoimmune syndrome (Selmi and others 2006). Among various interferon-inducible products are the PYHIN family proteins, which consists of a group of evolutionary related human and mouse proteins that have an N-terminal PYD domain and 1 or 2 partially conserved 200 amino acid long C-terminal HIN domain. Two members of the PYHIN family, namely AIM2 and interferon-inducible-16 (IFI16), have been shown to bind to and function as pattern recognition receptors (PRR) of virus-derived intracellular DNA, acting as foreign DNA sensor (Alnemri 2010; Brennan and Bowie 2010; Cristea and others 2010; Unterholzner and others 2010; Kerur and others 2011; Schattgen and Fitzgerald 2011; Unterholzner and Bowie 2011; Cridland and others 2012; Gariano and others 2012; Ratsimandresy and others 2013). These proteins display multifaceted activity due to their ability to bind to various target proteins (eg, transcription factors, signaling proteins, and tumor suppressor proteins) and modulate different cell functions. Increasing evidence supports a role for these proteins as regulators of various cell functions, including proliferation, differentiation, apoptosis, senescence, and inflammasome assembly (Xin and others 2003; Asefa and others 2004; Zhang and others 2007; Choubey and others 2008; Gariglio and others 2011; Veeranki and Choubey 2012).
The IFI16 protein is more specifically a nuclear phosphoprotein that is limited to vascular endothelial cells, keratinocytes, and hematopoietic cells (Gariglio and others 2002). IFI16 acts as an inducer of proinflammatory molecules (eg, ICAM-1, RANTES, and CCL20) and apoptosis when overexpressed in endothelial cells, supporting its role in the initial steps of the inflammatory processes that precede the onset of autoimmunity (Gugliesi and others 2005, 2010; Caposio and others 2007; Sponza and others 2009). In systemic autoimmune diseases, IFI16 protein is also a target for autoantibodies, as the presence of anti-IFI16 autoantibodies has been demonstrated in the sera of patients affected by SLE, SSc, and SjS (Mondini and others 2006, 2007; Costa and others 2011b; Agmon-Levin and others 2012; Rekvig and others 2012). This leads to an understanding that nuclear IFI16 can be mislocalized in the extracellular space inducing the production of specific autoantibodies. Recently, it has been demonstrated in vitro that the IFI16 protein can be induced to appear in the cytoplasm under conditions of UV-B-induced cell injury in normal keratinocytes and then released in the culture media (Costa and others 2011a).
In another scenario, IFI16, that acts as restriction factor against HCMV replication, has been found to be entrapped in egressing virions and exits the infected cells (Dell'Oste and others 2014). Moreover, it has been demonstrated that IFI16 is also mislocalized in KSHV latently infected cells, packaged in exosomes and released in the extracellular milieu (Singh and others 2013). However, since it was previously thought to be restricted to the intracellular environment, and in particular to the nucleus, all the recognized biological activities of IFI16 have been studied within the intracellular space (Schattgen and Fitzgerald 2011; Cridland and others 2012). In our previous studies, we have already shown that extracellular IFI16 can impair endothelial cell biological functions such as angiogenesis and migration. Such detrimental effects could be inhibited by masking the PYD domain of IFI16. Finally, we have also demonstrated that the extracellular IFI16 protein bound different cell lines with variable affinity, and it was later internalized and degraded (Gugliesi and others 2013).
In this article, we have further investigated the molecular activity of extracellular IFI16 protein on endothelial cells. Our novel findings describe the cytokine-stimulating activity of recombinant IFI16 (rIFI16) on primary endothelial cells, which can result in production and secretion of proinflammatory cytokines such as IL-6, IL-8, CCL2, CCL5, and CCL20. These results show a new function for circulating IFI16 at the endothelial interface and suggest a role for IFI16 in endothelial cell activation and injury in systemic inflammation. Overall, our article describes the proinflammatory behavior of extracellular IFI16 protein as one of the damage-associated molecular patterns (DAMPs) in autoimmune diseases.
Materials and Methods
Cell cultures
Primary human umbilical vein endothelial cells (HUVECs), pooled from multiple donors and cryopreserved at the end of the primary culture, were grown as described (Gugliesi and others 2010). Briefly, we used 0.2% gelatin-coated base in the presence of endothelial growth medium (EGM-2; Lonza-Milan) containing 2% fetal bovine serum, human recombinant vascular endothelial growth factor (rVEGF), basic fibroblast growth factor, human epidermal growth factor, IGF-1, hydrocortisone, ascorbic acid, heparin, gentamycin, and amphotericin B including 1% penicillin-streptomycin solution (Sigma-Aldrich), which we describe as complete EGM-2. All experiments were performed with low passage cells at 37°C and 5% CO2. Unless specified, HUVEC were grown in complete EGM-2, while 24 h before stimulation, they were grown in VEGF-deprived EGM-2.
Recombinant proteins
The entire coding sequence of the b-isoform of human IFI16 was subcloned into the pET30a expression vector (Novagen) containing an N-terminal histidine tag. Protein expression and nickel-affinity purification, followed by fast protein liquid chromatography (FPLC), were performed according to standard procedures. The purity of the proteins was assessed by 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis. The FPLC purified protein was then processed with Toxin Eraser™ Endotoxin Removal Kit (Genscript), while the lipopolysaccharide (LPS) concentration of the processed product was measured using Toxin Sensor Chromogenic LAL Endotoxin Assay Kit, which was as low as 0.05 EU/mL. The final purified rIFI16 was stored at −80°C in endotoxin-free vials. A preparation of BSA (Sigma Aldrich) was used as “mock” or “mock control” in all the stimulation experiments and was similarly prepared as rIFI16, briefly passed through the nickel-affinity column, LPS removal kit, and finally resuspended in rIFI16 buffer.
rIFI16 treatment and quantitative real-time polymerase chain reaction
For stimulation experiments, HUVEC were seeded (300,000 cells/well), grown overnight in 6-well plate, and stimulated with 50 μg/mL endotoxin-free (0.05 EU/mL) rIFI16 protein or mock. After stimulation, supernatants were sampled and stored at −80°C until further use for Cytokine ELISA, HUVEC were trypsinized and mRNA was extracted using TRI Reagent® (Sigma-Aldrich) as described in the manufacturer's protocol. The resulting mRNA was treated with DNase I Amplification Grade kit (Sigma-Aldrich) as instructed. Later, 1 μg of mRNA was used as a template first strand for cDNA synthesis using ImProm-II™ Reverse Transcription System (Promega) by following the manufacturer's protocol, while all polymerase chain reaction (PCR) reactions were carried out on C1000 Thermal Cycler (Bio-Rad). A random mRNA sample was kept as without reverse transcriptase (RT−) to assess the presence of contaminating genomic DNA in the preparation. The quantitative real-time PCR analyses were performed using CFX96 Real-Time PCR Detection System (Bio-Rad) with SsoAdvanced™ Universal SYBR® Green Supermix (Bio-Rad) and amplification conditions as instructed in the manufacturer's protocol, up to 40 cycles of PCR. Primer sequences are summarized in Supplementary Table S1 (Supplementary Data are available online at www.liebertpub.com/jir). The relative normalized expression after stimulation as compared to control was calculated as Fold Change=2−Δ(ΔCT) where ΔCT=CTtarget − CTGAPDH and Δ(ΔCT)=ΔCTstimulated − ΔCTcontrol.
Cytokine ELISA
The cytokines secreted in culture supernatants were analyzed using Single Analyte Human ELISA kits for IL-6, IL-8, CCL2 (Life Technologies), CCL5, and CCL20 (R&D Systems) according to the manufacturer's instructions. All absorbance readings were measured at 450 nM using a Victor X4 Multilabel Plate Reader (Perkin Elmer).
Flow cytometry
HUVEC were gently harvested by cell scrapper, washed twice in staining buffer (1% fetal calf serum, 0.01% NaN3 in phosphate buffered saline), and incubated with ICAM1-FITC (A15748; Life Technologies), VCAM1-PE (A16391; Life Technologies), and TLR4-PE (A18433; Life Technologies) antibodies separately at 4°C for 1 h. Later, the cells were washed twice again, resuspended in staining buffer, and analyzed by BD FACSCalibur. All analysis was performed using BD CellQuest Pro software.
siRNA transfection, antibodies, and inhibitors
FlexiTube siRNA for MyD88, Control siRNA, and HyPerFect transfection reagent were purchased from Qiagen and gene knockdown was performed according to the manufacturer's protocol. Transfection protocol was optimized for HUVEC to avoid cell viability issues. Antibodies used for western blots were p44/42 MAPK (#9102; Cell Signaling), p-p44/42 (#9101; Cell Signaling), p38 MAPK (#9212; Cell Signaling), p-p38 MAPK (#4511; Cell Signaling), JNK (#9258; Cell Signaling), pJNK (#9251; Cell Signaling), AKT (05-591; Merck Millipore), pAKT-Thr308 (#4056; Cell Signaling), pAKT-Ser473 (P4112; Sigma Aldrich), NFkB p65 (orb11118; Biorybt), NFkB p-p65 (#3036; Cell Signaling), p44/42 inhibitor-FR180204 (Sigma Aldrich), and p38 inhibitor-SB203580 (Invivogen).
Western blotting
Briefly, equal amount of cell extracts were fractionated by electrophoresis on SDS-polyacrylamide gels and transferred to Immobilon-P membranes (Millipore). After blocking in 5% nonfat milk, membranes were incubated overnight at 4°C with the appropriate primary antibodies. Membranes were then washed and incubated for 1 h at room temperature with secondary antibodies. Proteins were detected using an enhanced chemiluminescence detection kit (Thermo Scientific). Images were acquired and densitometry of the bands was performed using Quantity One software (version 4.6.9; Bio-Rad). Densitometry values were normalized using corresponding loading controls.
NF-κB immunofluorescence
Immunofluorescence was performed after 0, 4, and 8 h of rIFI16 treatment using 1:1,000 NF-κB p65 monoclonal antibody (F-6) sc-8008 (Santa Cruz Biotechnology, Inc.) as primary and 1:500 Alexa488-anti-mouse (GE Healthcare) as secondary antibody. Briefly, after treatment the cells were fixed with 2% paraformaldehyde (Sigma-Aldrich) for 20 min at 4°C, permeabilized with 0.5% Triton X-100 (Sigma-Aldrich) for 30 min at room temperature, incubated with primary antibody for 2 h at 37°C in a moist chamber, and incubated with secondary antibody and nuclear stain 1:500 TO-PRO-3 (Life-Technologies) for 1 h at room temperature. The glass cover slips were mounted using ProLong® Gold Antifade Reagent (Invitrogen) and observed under Leica DM-IRE2 Confocal Microscope.
Statistical analysis
All statistical tests were performed using GraphPad Prism version 6.00 for Windows (GraphPad). To test significance of rIFI16 treatment on endothelial cells by real time PCR, Cytokine ELISA, 2-way analysis of variance with Turkey's multiple comparison test was used. Significance of densitometric analysis was calculated using unpaired t-test. All data were plotted in histograms, bar graphs using mean±SD.
Results
rIFI16 induces time- and dose-dependent production of proinflammatory cytokines
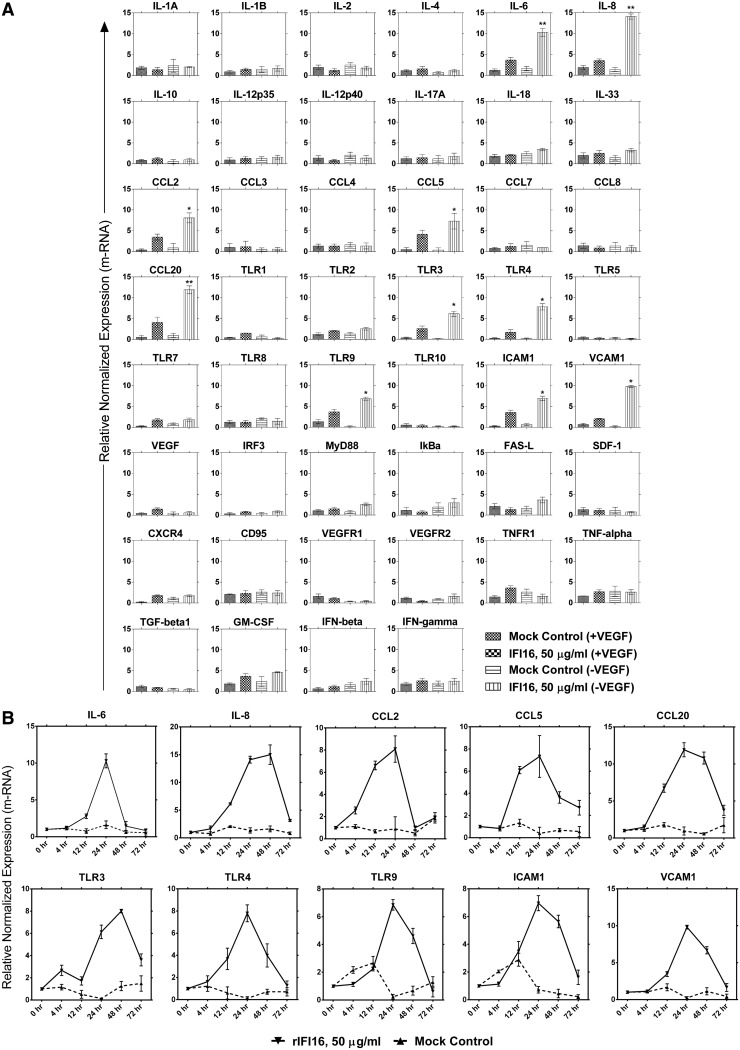
We specifically designed a set of SYBR green primers for real-time PCR (Supplementary Table S1) to identify the cytokine modulation pattern in rIFI16 stimulated endothelial cells. HUVEC were incubated with 50 μg/mL of endotoxin-free (<0.05 EU/mL) rIFI16 protein or mock control in presence or absence of VEGF in EGM-2 medium. After 24 h treatment, the relative normalized expression of mRNA was observed as shown in Fig. 1A. In a comparison between HUVEC grown with or without VEGF, higher gene modulation was identified in VEGF-depleted cells. Out of all the modulated genes, IL-6, IL-8, CCL2, CCL5, CCL20, TLR3, TLR4, TLR9, ICAM1, and VCAM1, displayed much significant upregulation in VEGF-depleted HUVEC. VEGF depletion before stimulation allows cell cycle synchronization to obtain equal response from different cells in a population. Time course experiments showed that IL-6, IL-8, CCL2, CCL5, and CCL20 mRNAs started to increase at 4 h and peaked between 12 and 48 h respectively (Fig. 1B). In contrast, the expression of inflammatory Toll-like receptors rose at later time points (24–48 h). Expression of cellular adhesion molecules like ICAM1 and VCAM1 started to increase at 4 h, and peaked at 24 h. Overall, our data indicated that inflammatory cytokines, chemokines, and adhesion molecules were expressed at an early stage followed by inflammatory receptors including TLR3, TLR4, and TLR9.
FIG. 1.
mRNA expression profile of proinflammatory cytokines in rIFI16-stimulated HUVEC. (A) Bar graphs representing folds change mRNA expression by HUVEC grown in presence/absence of VEGF growth factor in EGM-2, stimulated for 24 h with either rIFI16 (50 μg/mL) or mock control. (B) Solid and dotted line histograms representing fold change time-dependent mRNA expression by HUVEC grown in VEGF-depleted EGM-2, stimulated with either rIFI16 (50 μg/mL) or mock control for 0, 4, 12, 24, 48, and 72 h. Each real time PCR reaction was performed with Bio-Rad CFX96 and relative normalized expression of mRNA was calculated by the comparative Ct method. All experiments were performed in triplicates and the bar graphs/histograms represent (mean±SD) 6 independent experiments. All experimental data were processed by GraphPad Prism 6.01 software that was used to plot histograms and calculate statistical significance by unpaired t-test. *P<0.05; **P<0.01. HUVEC, human umbilical vein endothelial cell; rIFI16, recombinant interferon-inducible-16; VEGF, vascular endothelial growth factor.
To confirm that mRNA stimulation could be extended at protein level, we performed ELISA assay for each cytokine upregulated at the mRNA level. HUVEC were incubated with increasing concentrations of rIFI16 ranging from 1 μg/mL to 75 μg/mL, or mock control in VEGF-depleted EGM-2 for 24 h. As shown in Fig. 2A, for all of the cytokines analyzed, their concentration steadily increased with rIFI16 dosage. Consequently, we finalized 50 μg/mL of rIFI16 to test the cytokine stimulating properties in all the subsequent experiments. To observe the time-dependent cytokine expression profile, we tested supernatants from rIFI16 or mock control stimulated HUVEC at different time points. As shown in Fig. 2B, at 4 h time point, IL-6 (P<0.001) and IL-8 (P<0.001) achieved statistically significant levels in the supernatant as compared to corresponding mock-treated samples. In addition, the presence of CC chemokines such as CCL2 (P<0.01), CCL5 (P<0.05), and CCL20 (P<0.01) was also detected with lower significance. In the later time points, all the cytokines appeared to have increased in the supernatants due to timely accumulation. The secretion of IL-6 and IL-8 overall peaked by 12 h time point (P<0.001), while minimum increase was observed in the later stages. Other chemokines namely CCL2, CCL5, and CCL20 were strongly secreted starting from 8 h and peaked by 24 h time points, while almost steady levels in the later time points were observed. Consistent with the data obtained at the mRNA levels, all the cytokines analyzed by ELISA were significantly upregulated (P<0.001) at their peak time points.
FIG. 2.
Dose- and time-dependent secretion of proinflammatory cytokines upon rIFI16 treatment. (A) Dose-dependent secretion of multiple cytokines after rIFI16 treatment. Interleaved histograms indicating dose-dependent folds change in cytokine secretion by HUVEC stimulated with increasing concentration of rIFI16 (0, 1, 10, 25, 50, and 75 μg/mL respectively) for 24 h. (B) Time-dependent secretion of multiple cytokines after rIFI16 treatment. Interleaved histograms indicating time-dependent folds change in cytokine secretion by HUVEC stimulated with rIFI16 (50 μg/mL) for 0, 4, 8, 12, 24, and 48 h. All cells were grown in VEGF-depleted EGM-2 and ELISA was performed in triplicates using single analyte ELISA kits (Human) by Life Technologies. Each histogram represent (mean±SD) 6 independent experiments. All experimental data were processed by GraphPad Prism 6.01 software that was used to plot histograms and calculate statistical significance by 2-way ANOVA. *P<0.05; **P<0.01; ***P<0.001. ANOVA, analysis of variance.
LPS synergizes with rIFI16 through MyD88-dependent pathway for proinflammatory cytokine induction
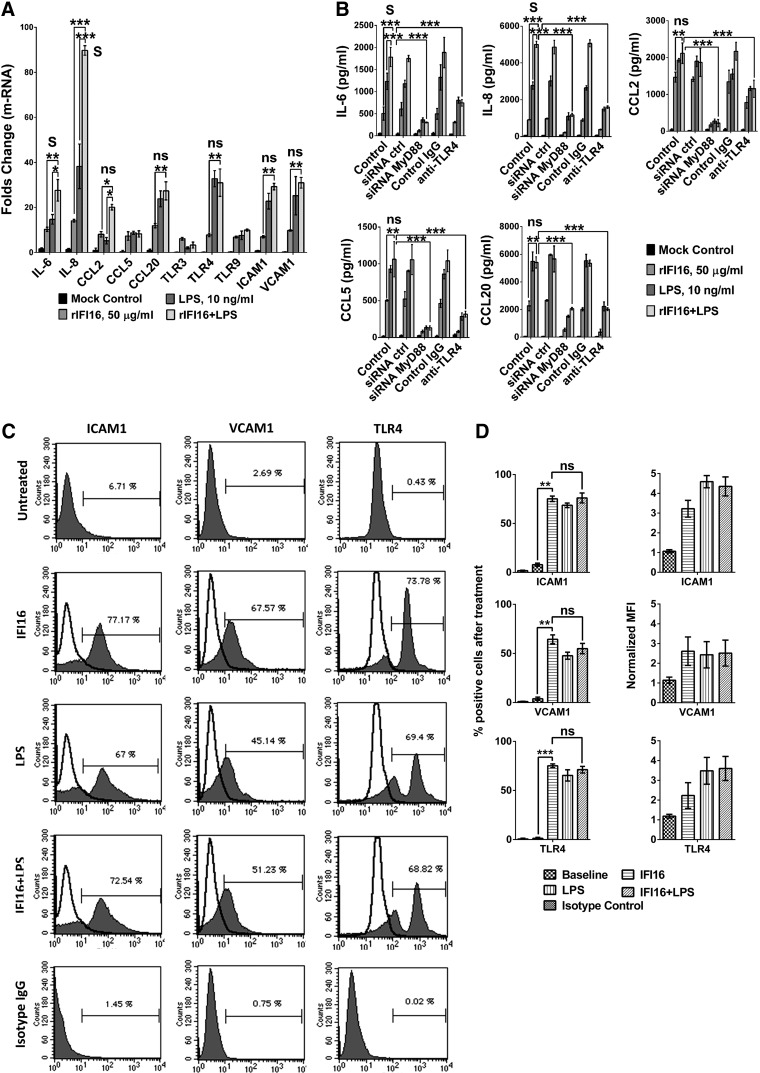
LPS alone is a potent inducer of inflammatory cytokines and by its interaction with TLR4 it drives activation of numerous transcription factors like NF-κB, IRF3, and others (Lu and others 2008). Here, we tested the stimulatory properties of rIFI16 in the presence of 10 ng/mL LPS preincubated overnight at room temperature to facilitate LPS-protein complexing. Twenty-four hours after rIFI16/LPS treatment, the mRNA expression profile of the set of positive genes from the previous experiments was analyzed by RT-PCR. As shown in Fig. 3A, the presence of LPS in rIFI16 preparation increased mRNA expression of IL-6, IL-8, CCL2, CCL20, TLR4, ICAM1, and VCAM1 as compared to rIFI16 alone while no significant change was observed in CCL5, TLR3, and TLR9. A comparison between mRNA expression levels by rIFI16, LPS, or rIFI16+LPS stimulation revealed synergistic increase in IL-6 and IL-8, but not the other cytokines. To corroborate these data, we tested the supernatants from the same 24 h treatment experiments by ELISA. As shown in Fig. 3B, LPS+IFI16 increased IL-6 and IL-8 production by 2-fold as compared to LPS alone and 3 to 5-fold when compared to rIFI16 alone. By contrast, secretion of CCL5, CCL20, and CCL2 was not upregulated by the combined treatment as compared to LPS or IFI16 alone.
FIG. 3.
rIFI16 synergizes with lipopolysaccharide (LPS) to increase proinflammatory cytokine expression through MyD88-dependent pathways. (A) HUVEC were grown in VEGF-depleted EGM-2 and stimulated for 24 h with either mock control, rIFI16 (50 μg/mL), LPS (10 ng/mL), or rIFI16+LPS mixture. Each RT-PCR reaction was performed with Bio-Rad CFX96 and relative normalized expression of mRNA was calculated using comparative Ct method. (B) MyD88/TLR4 silencing/neutralization inhibits rIFI16 cytokine stimulating activity and LPS synergy. HUVEC were stimulated for 24 h with mock control, rIFI16 (50 μg/mL), LPS (10 ng/mL), and rIFI16+LPS in the presence of no siRNA/antibodies (Control dataset), siRNA control, siRNA MyD88, Control IgG, or anti-TLR antibodies. (C) Flow cytometry histograms showing expression of ICAM1, VCAM1, and TLR4 in HUVEC when stimulated with mock control, rIFI16 (50 μg/mL), or LPS (10 ng/mL). One representative example from 6 independent experiments is shown. The shaded histograms represent binding of anti-ICAM1, anti-VCAM1, and anti-TLR4 antibodies and open histograms represent corresponding untreated controls. Isotype controls for each antibody were used in parallel to identify unspecific staining. (D) Histograms representing% positive cells after treatment and normalized mean fluorescence intensity as compared to isotype controls, plotted as mean±SD for ICAM1, VCAM1, and TLR4 from 6 independent experiments. All experimental data were processed by GraphPad Prism 6.01 software that was used to plot histograms and calculate statistical significance by 2-way ANOVA. *P<0.05; **P<0.01; ***P<0.001; s, synergy; ns, no synergy.
Next, to investigate the involvement of TLR pathways in rIFI16-mediated cytokine stimulation, we treated MyD88-silenced or TLR4-neutralized HUVEC with mock control, rIFI16, LPS, or rIFI16+LPS and 24 h later supernatants were analyzed for cytokines production. Overall, 4 to 6-fold decrease in the levels of IL-6, IL-8, CCL2, CCL5, and CCL20 was observed in MyD88-silenced cells as shown in Fig. 3B. In parallel experiments using TLR4 neutralizing antibodies, 2–3-fold decrease in the production of such cytokines was observed (Fig. 3B).
The upregulation of adhesion molecules like ICAM1, VCAM1, and inflammatory TLR4 receptor in response to external stimuli is a sensitive marker of endothelial cell activation and inflammation (Szmitko and others 2003). Here, we stimulated HUVEC with mock control, rIFI16, LPS, or combined for 24 h and the expression of ICAM1, VCAM1, and inflammatory TLR4 receptor was analyzed with flow cytometry, assessing the percentage positive cells and mean fluorescence intensity after treatment. We observed that the baseline expression of ICAM1 and VCAM1 before treatment was very low, while the cells were already expressing TLR4 before treatment. rIFI16 or LPS treatment per se similarly increased percentage of positive cells expressing ICAM1 (12 and 10-fold respectively), VCAM1 (25 and 16-fold respectively), and TLR4 (170 and 160-fold respectively) while no significant increase was detected with the combined treatment (Fig. 3C, D).
IL-6 and IL-8 induction by rIFI16 treatment requires differential activation of p38, p44/42, and p65
The activation of intracellular kinases is a very early event in response to inflammatory stimuli such as TNF-α and LPS and plays a central role in endothelial cell activation (Leonard and others 1999; Leong and Karsan 2000; Hoffmann and others 2002; Guijarro-Munoz and others 2014). To check for the time-dependent presence of active phosphorylated forms of pathway intermediates, western blot analysis was performed for NF-κB p65, p44/42, p38, and JNK in rIFI16-stimulated endothelial cells. As shown in Fig. 4A the phosphorylated form of p44/42 did not appear in the first 4 h of treatment, while it was present starting from 4 to 6 h. NF-κB p65 was seen evident starting from 1 h and peaked by 2 h, which was later downregulated. On the other hand, phospho-p38 showed immediate activation at 30 min time point. Moreover, JNK was not activated by rIFI16 treatment. The statistical significance for the expression of phosphoactive form of the above intermediates was calculated with densitometry by normalizing with corresponding basal levels of p44/42, p38, JNK, and NF-κB p65 respectively, as shown in Fig. 4B.
FIG. 4.
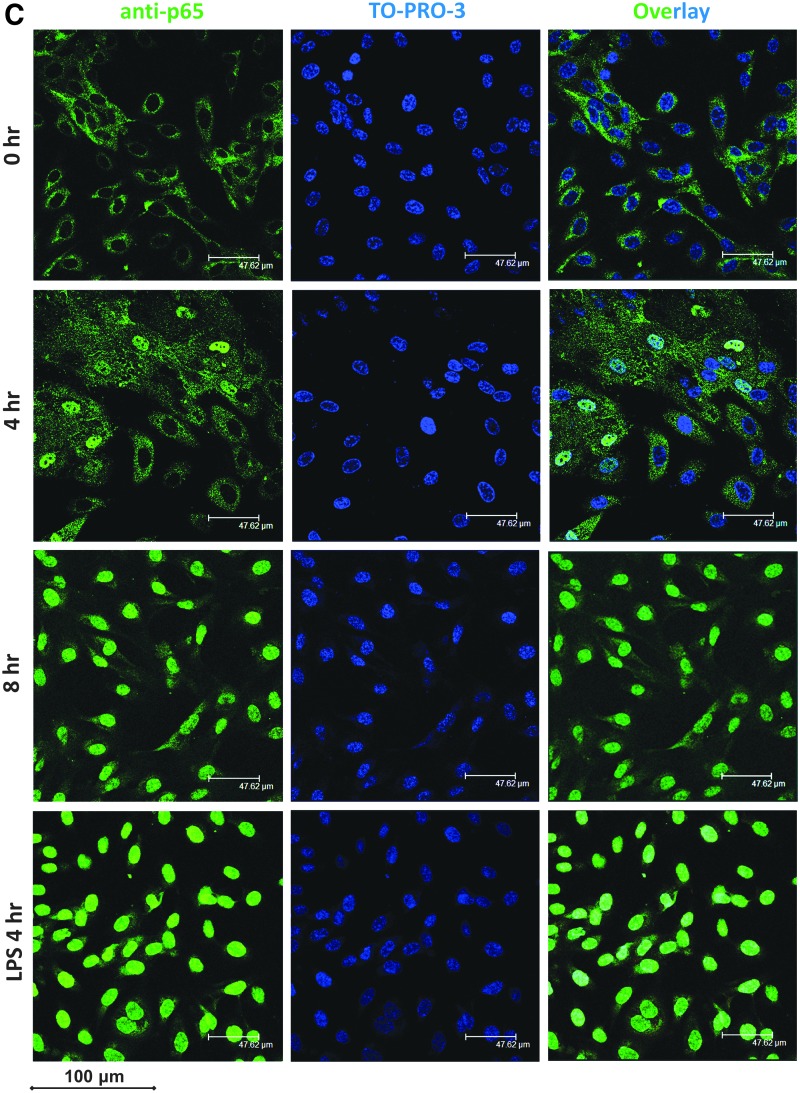
Differential activation of p44/42, p38, JNK, and NF-κB kinases in rIFI16-stimulated HUVEC. HUVEC were stimulated with rIFI16 (50 μg/mL) and the phosphoactive forms of (A) p44/42, p38, JNK, and NF-κB were detected by western blotting. One representative blot of 3 independent experiments is shown. Densitometric analysis predicting fold change in expression (mean±SD) from 3 independent experiments is represented by (B) corresponding histograms. Western blots of phosphoactive forms were stripped and incubated with antibodies against basal forms for loading control and densitometry normalization. All western blot images were acquired using VersaDoc 3000 equipment and images were exported using Quantity One 4.6.9 (Bio-Rad). Protein bands were cropped and compiled using Microsoft PowerPoint 2013 software. All experimental data were processed by GraphPad Prism 6.01 software that was used to plot histograms and calculate statistical significance by unpaired t-test. (C) HUVEC were stimulated with rIFI16 (50 μg/mL) for 0, 4, and 8 h and with LPS (10 ng/mL) for 4 h. Immunofluorescence was performed using 1:1000 anti-p65 polyclonal antibody (sc-109) and 1:500 TO-PRO-3 (Life Technologies) as nuclear counter stain. One representative image from 6 independent experiments is shown. *P<0.05.
The activation of NF-κB is an important event in the propagation of inflammation, since these transcription factors regulate the production of different proinflammatory cytokines (Tak and Firestein 2001; Lawrence 2009). After confirming NF-κB p65 phosphorylation by western blotting, we tested the proinflammatory activity of rIFI16 by analyzing the nuclear translocation of p65 subunit of NF-κB. As shown in Fig. 4C, in untreated cells, the p65 subunit was abundantly located in the cytoplasm, while in rIFI16-treated cells it started to migrate inside the nucleus in some cells and then, by 8 h, most of the cells were having complete p65 subunit translocated into the nucleus, confirming the activation of NF-κB. LPS (10 ng/mL), used as positive control, induced the nuclear translocation of p65 domain of NF-κB within the first 4 h.
Activation of p65 is independent and p44/42 is dependent on p38-mediated IL-6 and IL-8 secretion
To identify the role of p38 and p44/42 MAPK in the downstream signaling of rIFI16, cells were treated with the specific inhibitors SB203580 (p38 inhibitor) and FR180204 (p44/42 MAPK inhibitor) during 24 h rIFI16 stimulation and then production of IL-6 and IL-8 was measured by ELISA. As shown in Fig. 5A and B, SB203580 significantly downregulated the secretion of IL-6 (P<0.01) and IL-8 (P<0.01) respectively, while FR180204 only decreased IL-8 secretion by minimal amount (P<0.05). The combination of these inhibitors blocked these cytokines as SB203580 alone, which explains that p44/42 may not be directly involved in the secretion of first wave of cytokines such as IL-6 and IL-8.
FIG. 5.
Phosphorylation of NF-κB is independent and p44/42 is dependent on P-p38-mediated cytokine secretion. HUVEC were stimulated with rIFI16 (50 μg/mL) for 24 h in the presence of SB203580 (p38 inhibitor), FR180204 (p44/42 MAPK inhibitor), or both and the secretion of (A) IL-6 and (B) IL-8 was measured by ELISA. All experiments were carried out in triplicates and bar diagrams represent the levels of IL-6 and IL-8 (mean±SD) from 3 independent experiments. Western blots for (C) Phospho-p44/42 and (E) Phospho-NF-κB show the activation kinetics in the presence or absence of SB203580 in rIFI16-stimulated HUVEC. One representative blot from 3 independent experiments is shown. Histograms from panel (D) and (F) represent fold change in expression (mean±SD) calculated from 3 independent experiments by western blot densitometry study. Western blots of P-p44/42 and P-NF-κB were stripped and incubated with antibodies against basal p44/42 or NF-κB for loading control and densitometry normalization. Western blot images were acquired using VersaDoc 3000 equipment and images were exported using Quantity One 4.6.9 (Bio-Rad). Protein bands were cropped and compiled using Microsoft PowerPoint 2013 software. All experimental data were processed by GraphPad Prism 6.01 software that was used to plot histograms and calculate statistical significance by unpaired t-test. *P<0.05; **P<0.01; ns, not significant.
Next, to distinguish between the dependency of MAPK and NF-κB pathway intermediates in the same experimental setting, we used p38 inhibitor (SB203580) alone. As shown in Fig. 5C and D, SB203580 suppressed the late (4 to 6 h) activation of p44/42, but it did not inhibit the activation of p65 (Fig. 5E, F). These results are consistent with the lower concentrations of IL-6 and IL-8 observed even after p38 inhibition. Overall, our findings demonstrate that the late activation of p44/42 was dependent on the early activation of p38, while the activation of p65 was independent of p38 activation/phosphorylation.
Discussion
The IFI16 protein, a member of the PHYIN (p200) family, can be mislocalized in the cytoplasm and then released in the extracellular milieu as a consequence of several stimuli, including herpesvirus infection or UVB exposure (Costa and others 2011a; Kerur and others 2011; Gugliesi and others 2013; Dell'Oste and others 2014; Kivity and others 2014). The extracellular leaked IFI16 protein binds to high affinity sites on the plasma membrane of primary endothelial cells, causing detrimental changes in their biological activity, including angiogenesis and cellular migration (Gugliesi and others 2013).
In this study, we demonstrate that extracellular IFI16 functions as a DAMP propagating “danger signal” that induces a proinflammatory phenotype in endothelial cells in vitro characterized by upregulation of leukocyte adhesion molecules (ICAM-1 and VCAM-1), secretion of the neutrophil and monocyte stimulatory chemokines (IL-6, IL-8, CCL2, CCL5, and CCL20,), and enhanced expression of surface TLR4. The expression of these cytokines was linked with the activation of multiple signaling pathways downstream involving the activation of MAPK and NF-κB intermediates. Our data explain p38 MAPK as a key downstream effector of rIFI16 stimulation as its activity is required for activation of p44/42 MAPK and cytokine secretion. Moreover, p38 activation was an early event while p42/44 MAPK activity peaked between 4 and 5 h when IFI16-induced cytokines reached significant amounts in cell supernatant (Figs. 2 and 5). These data led us to speculate that the bulk of p42/44 MAPK activity is not directly due to IFI16 but to an autocrine/paracrine loop resulting from cytokine release by IFI16-stimulated cells. Conversely, rIFI16 stimulation induced the phosphorylation of p65 NF-κB subunit in a manner that was p38 independent, probably justifying the low amounts of cytokines released also in presence of p38 inhibitors.
The environment created by the IFI16 protein is similar to the functioning of different DAMPs such as HMGB1 and Heat Shock Proteins (Bianchi 2007; Piccinini and Midwood 2010; Castiglioni and others 2011; Rosin and Okusa 2011; Said-Sadier and Ojcius 2012; Moudgil and others 2013; Pisetsky 2013; Schiopu and Cotoi 2013; Pouwels and others 2014; Tsai and others 2014). A majority of DAMPs have shown to alarm the immune system by the secretion of potent inflammatory cytokines such as IL-6, and TNF-α, which in turn stimulate different pathways for the secretion of numerous other cytokines (Fiuza and others 2003; Harris and others 2012). In our case, we also tested the upregulation of TNF-α mRNA in IFI16-stimulated cells, but no evidence of its upregulation was found. Previous gene array analysis of HUVECs overexpressing IFI16 has revealed an increased expression of genes involved in inflammation and apoptosis. Functional analysis of the ICAM-1 promoter by deletion- or site-specific mutation demonstrated that NF-kB is the main mediator of IFI16-driven gene induction (Caposio and others 2007). Consistent with these studies, protein array analysis of the IFI16 secretome showed an increased production of chemokines, cytokines, and adhesion molecules responsible for leukocyte chemotaxis. Functional analysis of the promoter for CCL20, the chemokine responsible for leukocyte recruitment in the early steps of inflammation, by site-specific mutation demonstrated that NF-kB is the main mediator of CCL20 induction at the transcriptional level (Baggetta and others 2010). In both studies no induction of TNF-α was observed suggesting that the presence of NF-kB responsive elements is not sufficient for gene induction, but other regulatory mechanisms intervene to discriminate among gene promoters containing NF-kB responsive DNA motifs. Another common aspect of DAMPs stimulating chronic inflammation is contributed by their synergy with LPS (Qin and others 2009; Chakraborty and others 2013; He and others 2013). IFI16 alone was able to stimulate significant amounts of inflammatory cytokine expression, but in the presence of working concentrations of LPS (10 ng/mL), such activity was amplified, especially for IL-6 and IL-8 production.
The extracellular spillage of endogenous molecules can uncontrollably activate the immune system, while chronic stimulation could be a possible lead toward the development of an autoimmune syndrome (Toubi and Shoenfeld 2004; Molina and Shoenfeld 2005; Harris and others 2012). Proinflammatory cytokine secretion triggered by DAMPs and PAMPs is widely regulated by inflammatory TLR receptors, while MyD88 form a universal adapter protein, which transduces several downstream signaling pathways after TLR activation (Midwood and others 2009; Piccinini and Midwood 2010). The nuclear DNA sensor IFI16 behaves in similar DAMP-like manner when mislocalized from its nuclear compartment into the extracellular milieu. The activity of IFI16, LPS, and LPS complexed IFI16 was drastically reduced in MyD88 silenced endothelial cells, suggesting a role of MyD88-dependent signaling pathways in IFI16-mediated cytokine secretion, while such significant reduction in cytokine secretion was also observed in TLR4 neutralized cells, which further supports our data. These findings, together with the recently documented occurrence of the IFI16 protein in the sera of systemic autoimmune patients suggest a role for IFI16 as an alarmin driving the pathogenesis of inflammatory and autoimmune disease.
With this scenario in mind, it is becoming clear that the induction of specific anti-IFI16 autoantibodies in patients with systemic autoimmune diseases (Mondini and others 2007; Rekvig and others 2012) is triggered by the presence of IFI16 in the extracellular space with the contribution of its chronic inflammation stimulating activity. Inside the cell, IFI16 binds DNA in a sequence-independent manner and sense nonself DNA (Morrone and others 2014); this location is not fixed, however, and, with viral infection, injury, or death, it can translocate outside of the cell (Costa and others 2011a; Gariano and others 2012; Li and others 2012; Gugliesi and others 2013; Dell'Oste and others 2014). In the extracellular milieau, IFI16 can serve as a DAMP or alarmin to amplify inflammation and stimulate the innate immune system.
The mRNA expression data revealed that rIFI16 stimulation also upregulated TLR-9 in endothelial cells. However, the mRNA of TLR9-mediated cytokines, namely IL-12, TNF-α, and IFN-γ was not modulated, which suggests lack of TLR9 activation. It potentially shows that IFI16, which is mislocalized during viral infection in the extracellular milieu (Singh and others 2013; Dell'Oste and others 2014), is interacting with unknown cell membrane PRRs, which could lead to the upregulation of TLR9, just to alarm the innate immune system for the presence of PAMPs. In this direction, there is some evidence indicating that recombinant FITC-labeled IFI16 can be internalized by primary endothelial cells (Gugliesi and others 2013), which could also expose it to intracellular PRRs. Two very recent publications describe the presence of inflammasome component ASC and the inflammasome sensor NLP3 in the extracellular space and their function as danger signal that amplifies the inflammatory response both in vitro and in vivo (Baroja-Mazo and others 2014; Franklin and others 2014). These extracellular inflammasome components are internalized by macrophages to initiate caspase-1 secretome, promoting inflammation. IFI16 is another inflammasome sensor with a nuclear localization and a restricted expression in endothelial cells, keratinocytes, and some hemopoietic cells (Gariglio and others 2002). Moreover, the group of Bala Chandran recently demonstrated that the nuclear resident IFI16 protein is involved in the sensing of EBV and KSHV genome, which leads to the formation of the active IFI16–ASC–caspase-1 inflammasome and production of the cleaved form of proinflammatory IL-1b, IL-18, and IL-33 cytokines that are released together with IFI16 in the exosomes of the culture supernatant of cells harboring the viral genomes (Kerur and others 2011; Ansari and others 2013). In addition, our group has described that IFI16 is mislocalized and eventually found in the extracellular space and virions during lytic cytomegalovirus infection and keratinocytes exposure to UVB (Costa and others 2011a; Dell'Oste and others 2014).
Consistent with these observations, this study reveals previously unappreciated extracellular activities of IFI16 that lead to new cell to cell communication mechanism and chronic amplification of the inflammatory signal. Since the intracellular IFI16 was already reported as a regulator of endothelial proinflammatory activity, our novel findings strengthen its role in the initiation, propagation, and maintenance of the inflammatory responses in the endothelium inside and outside the cell during infection or injury. These in vitro data are fully consistent with the available in vivo data on IFI16 presence as both circulating protein and specific autoantibodies in the sera of patients affected by systemic autoimmune diseases and with its role in the propagation of inflammation and development of autoimmune disorders. Another interesting aspect that emerges from this study is the synergistic action of IFI16 with LPS indicating its possible involvement also as promoter of inflammation in microorganism-induced tissue damage. This explains the emerging relevance of DNA sensors or other inflammasome components in the initiation and propagation of chronic inflammation, when leaked outside the cellular compartment due to injury, viral infection, or physical agents.
Supplementary Material
Acknowledgments
This study was supported by grant Ministero dell'Istruzione, dell'Università e della Ricerca-Progetti di Ricerca di Interesse Nazionale (MIUR-PRIN) 2012 to S.L. (2012SNMJRL); and the University of Turin in 2013 to S.L. and M.D.A. M.B. is a recipient of an international PhD fellowship in Innovative Biomedical Technologies (IBT) funded by Cariplo Foundation-Milan, Italy.
Authorship Contributions
S.L. and M.G. conceived, designed the experiments, and wrote the article. M.B., M.D.A., I.L.C., and V.C. performed the experiments and analyzed the data. G.B. and A.G. provided reagents and expertise for experiments involving MAPK and NF-κB.
Author Disclosure Statement
All authors declare no competing financial interests exist.
References
- Agmon-Levin N, Mosca M, Petri M, Shoenfeld Y. 2012. Systemic lupus erythematosus one disease or many? Autoimmun Rev 11(8):593–595 [DOI] [PubMed] [Google Scholar]
- Alnemri ES. 2010. Sensing cytoplasmic danger signals by the inflammasome. J Clin Immunol 30(4):512–519 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ansari MA, Singh VV, Dutta S, Veettil MV, Dutta D, Chikoti L, Lu J, Everly D, Chandran B. 2013. Constitutive interferon-inducible protein 16-inflammasome activation during Epstein-Barr virus latency I, II, and III in B and epithelial cells. J Virol 87(15):8606–8623 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Asefa B, Klarmann KD, Copeland NG, Gilbert DJ, Jenkins NA, Keller JR. 2004. The interferon-inducible p200 family of proteins: a perspective on their roles in cell cycle regulation and differentiation. Blood Cells Mol Dis 32(1):155–167 [DOI] [PubMed] [Google Scholar]
- Baggetta R, De Andrea M, Gariano GR, Mondini M, Ritta M, Caposio P, Cappello P, Giovarelli M, Gariglio M, Landolfo S. 2010. The interferon-inducible gene IFI16 secretome of endothelial cells drives the early steps of the inflammatory response. Eur J Immunol 40(8):2182–2189 [DOI] [PubMed] [Google Scholar]
- Baroja-Mazo A, Martin-Sanchez F, Gomez AI, Martinez CM, Amores-Iniesta J, Compan V, Barbera-Cremades M, Yague J, Ruiz-Ortiz E, Anton J, Bujan S, Couillin I, Brough D, Arostegui JI, Pelegrin P. 2014. The NLRP3 inflammasome is released as a particulate danger signal that amplifies the inflammatory response. Nat Immunol 15:738–748 [DOI] [PubMed] [Google Scholar]
- Bennett L, Palucka AK, Arce E, Cantrell V, Borvak J, Banchereau J, Pascual V. 2003. Interferon and granulopoiesis signatures in systemic lupus erythematosus blood. J Exp Med 197(6):711–723 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bianchi ME. 2007. DAMPs, PAMPs and alarmins: all we need to know about danger. J Leukoc Biol 81(1):1–5 [DOI] [PubMed] [Google Scholar]
- Brennan K, Bowie AG. 2010. Activation of host pattern recognition receptors by viruses. Curr Opin Microbiol 13(4):503–507 [DOI] [PubMed] [Google Scholar]
- Bronson PG, Chaivorapol C, Ortmann W, Behrens TW, Graham RR. 2012. The genetics of type I interferon in systemic lupus erythematosus. Curr Opin Immunol 24(5):530–537 [DOI] [PubMed] [Google Scholar]
- Caposio P, Gugliesi F, Zannetti C, Sponza S, Mondini M, Medico E, Hiscott J, Young HA, Gribaudo G, Gariglio M, Landolfo S. 2007. A novel role of the interferon-inducible protein IFI16 as inducer of proinflammatory molecules in endothelial cells. J Biol Chem 282(46):33515–33529 [DOI] [PubMed] [Google Scholar]
- Castiglioni A, Canti V, Rovere-Querini P, Manfredi AA. 2011. High-mobility group box 1 (HMGB1) as a master regulator of innate immunity. Cell Tissue Res 343(1):189–199 [DOI] [PubMed] [Google Scholar]
- Chakraborty R, Bhatt KH, Sodhi A. 2013. High mobility group box 1 protein synergizes with lipopolysaccharide and peptidoglycan for nitric oxide production in mouse peritoneal macrophages in vitro. Mol Immunol 54(1):48–57 [DOI] [PubMed] [Google Scholar]
- Choubey D, Deka R, Ho SM. 2008. Interferon-inducible IFI16 protein in human cancers and autoimmune diseases. Front Biosci 13:598–608 [DOI] [PubMed] [Google Scholar]
- Costa S, Borgogna C, Mondini M, De Andrea M, Meroni PL, Berti E, Gariglio M, Landolfo S. 2011a. Redistribution of the nuclear protein IFI16 into the cytoplasm of ultraviolet B-exposed keratinocytes as a mechanism of autoantigen processing. Br J Dermatol 164(2):282–290 [DOI] [PubMed] [Google Scholar]
- Costa S, Mondini M, Caneparo V, Afeltra A, Airo P, Bellisai F, Faggioli P, Gerli R, Lotzniker M, Meroni PL, Morozzi G, Radice A, Riccieri V, Scarsi M, Sebastiani GD, Sinico RA, Tincani A, Gariglio M, Landolfo S. 2011b. Detection of anti-IFI16 antibodies by ELISA: clinical and serological associations in systemic sclerosis. Rheumatology (Oxford) 50(4):674–681 [DOI] [PubMed] [Google Scholar]
- Cridland JA, Curley EZ, Wykes MN, Schroder K, Sweet MJ, Roberts TL, Ragan MA, Kassahn KS, Stacey KJ. 2012. The mammalian PYHIN gene family: phylogeny, evolution and expression. BMC Evol Biol 12:140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cristea IM, Moorman NJ, Terhune SS, Cuevas CD, O'Keefe ES, Rout MP, Chait BT, Shenk T. 2010. Human cytomegalovirus pUL83 stimulates activity of the viral immediate-early promoter through its interaction with the cellular IFI16 protein. J Virol 84(15):7803–7814 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dell'Oste V, Gatti D, Gugliesi F, De Andrea M, Bawadekar M, Lo Cigno I, Biolatti M, Vallino M, Marschall M, Gariglio M, Landolfo S. 2014. Innate nuclear sensor IFI16 translocates into the cytoplasm during the early stage of in vitro human cytomegalovirus infection and is entrapped in the egressing virions during the late stage. J Virol 88(12):6970–6982 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fiuza C, Bustin M, Talwar S, Tropea M, Gerstenberger E, Shelhamer JH, Suffredini AF. 2003. Inflammation-promoting activity of HMGB1 on human microvascular endothelial cells. Blood 101(7):2652–2660 [DOI] [PubMed] [Google Scholar]
- Forster S. 2012. Interferon signatures in immune disorders and disease. Immunol Cell Biol 90(5):520–527 [DOI] [PubMed] [Google Scholar]
- Franklin BS, Bossaller L, De Nardo D, Ratter JM, Stutz A, Engels G, Brenker C, Nordhoff M, Mirandola SR, Al-Amoudi A, Mangan MS, Zimmer S, Monks BG, Fricke M, Schmidt RE, Espevik T, Jones B, Jarnicki AG, Hansbro PM, Busto P, Marshak-Rothstein A, Hornemann S, Aguzzi A, Kastenmuller W, Latz E. 2014. The adaptor ASC has extracellular and ‘prionoid’ activities that propagate inflammation. Nat Immunol 15:727–737 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gariano GR, Dell'Oste V, Bronzini M, Gatti D, Luganini A, De Andrea M, Gribaudo G, Gariglio M, Landolfo S. 2012. The intracellular DNA sensor IFI16 gene acts as restriction factor for human cytomegalovirus replication. PLoS Pathog 8(1):e1002498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gariglio M, Azzimonti B, Pagano M, Palestro G, De Andrea M, Valente G, Voglino G, Navino L, Landolfo S. 2002. Immunohistochemical expression analysis of the human interferon-inducible gene IFI16, a member of the HIN200 family, not restricted to hematopoietic cells. J Interferon Cytokine Res 22(7):815–821 [DOI] [PubMed] [Google Scholar]
- Gariglio M, Mondini M, De Andrea M, Landolfo S. 2011. The multifaceted interferon-inducible p200 family proteins: from cell biology to human pathology. J Interferon Cytokine Res 31(1):159–172 [DOI] [PubMed] [Google Scholar]
- Gugliesi F, Bawadekar M, De Andrea M, Dell'Oste V, Caneparo V, Tincani A, Gariglio M, Landolfo S. 2013. Nuclear DNA sensor IFI16 as circulating protein in autoimmune diseases is a signal of damage that impairs endothelial cells through high-affinity membrane binding. PLoS One 8(5):e63045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gugliesi F, De Andrea M, Mondini M, Cappello P, Giovarelli M, Shoenfeld Y, Meroni P, Gariglio M, Landolfo S. 2010. The proapoptotic activity of the Interferon-inducible gene IFI16 provides new insights into its etiopathogenetic role in autoimmunity. J Autoimmun 35(2):114–123 [DOI] [PubMed] [Google Scholar]
- Gugliesi F, Mondini M, Ravera R, Robotti A, de Andrea M, Gribaudo G, Gariglio M, Landolfo S. 2005. Up-regulation of the interferon-inducible IFI16 gene by oxidative stress triggers p53 transcriptional activity in endothelial cells. J Leukoc Biol 77(5):820–829 [DOI] [PubMed] [Google Scholar]
- Guijarro-Munoz I, Compte M, Alvarez-Cienfuegos A, Alvarez-Vallina L, Sanz L. 2014. Lipopolysaccharide activates Toll-like receptor 4 (TLR4)-mediated NF-kappaB signaling pathway and proinflammatory response in human pericytes. J Biol Chem 289(4):2457–2468 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harris HE, Andersson U, Pisetsky DS. 2012. HMGB1: a multifunctional alarmin driving autoimmune and inflammatory disease. Nat Rev Rheumatol 8(4):195–202 [DOI] [PubMed] [Google Scholar]
- He ZW, Qin YH, Wang ZW, Chen Y, Shen Q, Dai SM. 2013. HMGB1 acts in synergy with lipopolysaccharide in activating rheumatoid synovial fibroblasts via p38 MAPK and NF-kappaB signaling pathways. Mediators Inflamm 2013:596716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Higgs BW, Liu Z, White B, Zhu W, White WI, Morehouse C, Brohawn P, Kiener PA, Richman L, Fiorentino D, Greenberg SA, Jallal B, Yao Y. 2011. Patients with systemic lupus erythematosus, myositis, rheumatoid arthritis and scleroderma share activation of a common type I interferon pathway. Ann Rheum Dis 70(11):2029–2036 [DOI] [PubMed] [Google Scholar]
- Hoffmann E, Dittrich-Breiholz O, Holtmann H, Kracht M. 2002. Multiple control of interleukin-8 gene expression. J Leukoc Biol 72(5):847–855 [PubMed] [Google Scholar]
- Kerur N, Veettil MV, Sharma-Walia N, Bottero V, Sadagopan S, Otageri P, Chandran B. 2011. IFI16 acts as a nuclear pathogen sensor to induce the inflammasome in response to Kaposi Sarcoma-associated herpesvirus infection. Cell Host Microbe 9(5):363–375 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kivity S, Arango MT, Ehrenfeld M, Tehori O, Shoenfeld Y, Anaya JM, Agmon-Levin N. 2014. Infection and autoimmunity in Sjogren's syndrome: a clinical study and comprehensive review. J Autoimmun 51:17–22 [DOI] [PubMed] [Google Scholar]
- Lawrence T. 2009. The nuclear factor NF-kappaB pathway in inflammation. Cold Spring Harb Perspect Biol 1(6):a001651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leonard M, Ryan MP, Watson AJ, Schramek H, Healy E. 1999. Role of MAP kinase pathways in mediating IL-6 production in human primary mesangial and proximal tubular cells. Kidney Int 56(4):1366–1377 [DOI] [PubMed] [Google Scholar]
- Leong KG, Karsan A. 2000. Signaling pathways mediated by tumor necrosis factor alpha. Histol Histopathol 15(4):1303–1325 [DOI] [PubMed] [Google Scholar]
- Li T, Diner BA, Chen J, Cristea IM. 2012. Acetylation modulates cellular distribution and DNA sensing ability of interferon-inducible protein IFI16. Proc Natl Acad Sci U S A 109(26):10558–10563 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu YC, Yeh WC, Ohashi PS. 2008. LPS/TLR4 signal transduction pathway. Cytokine 42(2):145–151 [DOI] [PubMed] [Google Scholar]
- Midwood KS, Piccinini AM, Sacre S. 2009. Targeting Toll-like receptors in autoimmunity. Curr Drug Targets 10(11):1139–1155 [DOI] [PubMed] [Google Scholar]
- Molina V, Shoenfeld Y. 2005. Infection, vaccines and other environmental triggers of autoimmunity. Autoimmunity 38(3):235–245 [DOI] [PubMed] [Google Scholar]
- Mondini M, Vidali M, Airo P, De Andrea M, Riboldi P, Meroni PL, Gariglio M, Landolfo S. 2007. Role of the interferon-inducible gene IFI16 in the etiopathogenesis of systemic autoimmune disorders. Ann N Y Acad Sci 1110:47–56 [DOI] [PubMed] [Google Scholar]
- Mondini M, Vidali M, De Andrea M, Azzimonti B, Airo P, D'Ambrosio R, Riboldi P, Meroni PL, Albano E, Shoenfeld Y, Gariglio M, Landolfo S. 2006. A novel autoantigen to differentiate limited cutaneous systemic sclerosis from diffuse cutaneous systemic sclerosis: the interferon-inducible gene IFI16. Arthritis Rheum 54(12):3939–3944 [DOI] [PubMed] [Google Scholar]
- Morrone SR, Wang T, Constantoulakis LM, Hooy RM, Delannoy MJ, Sohn J. 2014. Cooperative assembly of IFI16 filaments on dsDNA provides insights into host defense strategy. Proc Natl Acad Sci U S A 111(1):E62–E71 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moudgil KD, Thompson SJ, Geraci F, De Paepe B, Shoenfeld Y. 2013. Heat-shock proteins in autoimmunity. Autoimmune Dis 2013:621417. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piccinini AM, Midwood KS. 2010. DAMPening inflammation by modulating TLR signalling. Mediators Inflamm 2010:pii: [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pisetsky DS. 2013. The effectors of innate immunity: DAMPs, DAMEs, or DIMEs? Arthritis Res Ther 15(5):123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pouwels SD, Heijink IH, ten Hacken NH, Vandenabeele P, Krysko DV, Nawijn MC, van Oosterhout AJ. 2014. DAMPs activating innate and adaptive immune responses in COPD. Mucosal Immunol 7(2):215–226 [DOI] [PubMed] [Google Scholar]
- Qin YH, Dai SM, Tang GS, Zhang J, Ren D, Wang ZW, Shen Q. 2009. HMGB1 enhances the proinflammatory activity of lipopolysaccharide by promoting the phosphorylation of MAPK p38 through receptor for advanced glycation end products. J Immunol 183(10):6244–6250 [DOI] [PubMed] [Google Scholar]
- Ratsimandresy RA, Dorfleutner A, Stehlik C. 2013. An update on PYRIN domain-containing pattern recognition receptors: from immunity to pathology. Front Immunol 4:440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rekvig OP, Putterman C, Casu C, Gao HX, Ghirardello A, Mortensen ES, Tincani A, Doria A. 2012. Autoantibodies in lupus: culprits or passive bystanders? Autoimmun Rev 11(8):596–603 [DOI] [PubMed] [Google Scholar]
- Ronnblom L, Alm GV, Eloranta ML. 2011. The type I interferon system in the development of lupus. Semin Immunol 23(2):113–121 [DOI] [PubMed] [Google Scholar]
- Ronnblom L, Eloranta ML. 2013. The interferon signature in autoimmune diseases. Curr Opin Rheumatol 25(2):248–253 [DOI] [PubMed] [Google Scholar]
- Rosin DL, Okusa MD. 2011. Dangers within: DAMP responses to damage and cell death in kidney disease. J Am Soc Nephrol 22(3):416–425 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruiz-Riol M, Barnils Mdel P, Colobran Oriol R, Pla AS, Borras Serres FE, Lucas-Martin A, Martinez Caceres EM, Pujol-Borrell R. 2011. Analysis of the cumulative changes in Graves' disease thyroid glands points to IFN signature, plasmacytoid DCs and alternatively activated macrophages as chronicity determining factors. J Autoimmun 36(3–4):189–200 [DOI] [PubMed] [Google Scholar]
- Said-Sadier N, Ojcius DM. 2012. Alarmins, inflammasomes and immunity. Biomed J 35(6):437–449 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schattgen SA, Fitzgerald KA. 2011. The PYHIN protein family as mediators of host defenses. Immunol Rev 243(1):109–118 [DOI] [PubMed] [Google Scholar]
- Schiopu A, Cotoi OS. 2013. S100A8 and S100A9: DAMPs at the crossroads between innate immunity, traditional risk factors, and cardiovascular disease. Mediators Inflamm 2013:828354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Selmi C, Lleo A, Zuin M, Podda M, Rossaro L, Gershwin ME. 2006. Interferon alpha and its contribution to autoimmunity. Curr Opin Investig Drugs 7(5):451–456 [PubMed] [Google Scholar]
- Singh VV, Kerur N, Bottero V, Dutta S, Chakraborty S, Ansari MA, Paudel N, Chikoti L, Chandran B. 2013. Kaposi's sarcoma-associated herpesvirus latency in endothelial and B cells activates gamma interferon-inducible protein 16-mediated inflammasomes. J Virol 87(8):4417–4431 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sozzani S, Bosisio D, Scarsi M, Tincani A. 2010. Type I interferons in systemic autoimmunity. Autoimmunity 43(3):196–203 [DOI] [PubMed] [Google Scholar]
- Sponza S, De Andrea M, Mondini M, Gugliesi F, Gariglio M, Landolfo S. 2009. Role of the interferon-inducible IFI16 gene in the induction of ICAM-1 by TNF-alpha. Cell Immunol 257(1–2):55–60 [DOI] [PubMed] [Google Scholar]
- Szmitko PE, Wang CH, Weisel RD, de Almeida JR, Anderson TJ, Verma S. 2003. New markers of inflammation and endothelial cell activation: Part I. Circulation 108(16):1917–1923 [DOI] [PubMed] [Google Scholar]
- Tak PP, Firestein GS. 2001. NF-kappaB: a key role in inflammatory diseases. J Clin Invest 107(1):7–11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Toubi E, Shoenfeld Y. 2004. Toll-like receptors and their role in the development of autoimmune diseases. Autoimmunity 37(3):183–188 [DOI] [PubMed] [Google Scholar]
- Tsai SY, Segovia JA, Chang TH, Morris IR, Berton MT, Tessier PA, Tardif MR, Cesaro A, Bose S. 2014. DAMP molecule S100A9 acts as a molecular pattern to enhance inflammation during influenza A virus infection: role of DDX21-TRIF-TLR4-MyD88 pathway. PLoS Pathog 10(1):e1003848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Unterholzner L, Bowie AG. 2011. Innate DNA sensing moves to the nucleus. Cell Host Microbe 9(5):351–353 [DOI] [PubMed] [Google Scholar]
- Unterholzner L, Keating SE, Baran M, Horan KA, Jensen SB, Sharma S, Sirois CM, Jin T, Latz E, Xiao TS, Fitzgerald KA, Paludan SR, Bowie AG. 2010. IFI16 is an innate immune sensor for intracellular DNA. Nat Immunol 11(11):997–1004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Veeranki S, Choubey D. 2012. Interferon-inducible p200-family protein IFI16, an innate immune sensor for cytosolic and nuclear double-stranded DNA: regulation of subcellular localization. Mol Immunol 49(4):567–571 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xin H, Curry J, Johnstone RW, Nickoloff BJ, Choubey D. 2003. Role of IFI 16, a member of the interferon-inducible p200-protein family, in prostate epithelial cellular senescence. Oncogene 22(31):4831–4840 [DOI] [PubMed] [Google Scholar]
- Zhang Y, Howell RD, Alfonso DT, Yu J, Kong L, Wittig JC, Liu CJ. 2007. IFI16 inhibits tumorigenicity and cell proliferation of bone and cartilage tumor cells. Front Biosci 12:4855–4863 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.