Table 1. Data-collection and refinement statistics.
Values in parentheses are for the outer shell except where indicated otherwise.
| SERCACa2AMPPCP | SERCAVO3TNPATP | SsZntAAlF4 | |
|---|---|---|---|
| Data-collection and processing | |||
| X-ray energy (keV) | 6 | 6 | 6 |
| Collected frames | 761730 | 1371609 | 280281 |
| Diffraction hits† | 23016 (3.0) | 83934 (6.1) | 16358 (5.8) |
| Indexed frames‡ | 4069 (17.8) | 4910 (5.8) | 55 (0.3) |
| Space group | C2 | P4212 | P422 |
| a, b, c () | 162, 76.3, 151 | 268, 268, 114.5 | 58, 58, 320 |
| , , () | 90, 109.0, 90 | 90, 90, 90 | 90, 90, 90 |
| Highest resolution observed () | 2.8 | 4 | 4 |
| Resolution range for processing () | 59.92.8 (2.92.8) | 57.255.00 (5.185.00) | n.a.§ |
| Unique reflections | 42416 (3652) | 18601¶ (1813) | n.a.§ |
| Multiplicity | 17.3 (4.2) | 124 (117) | n.a.§ |
| I/(I) | 1.13 (0.39) | 2.03 (0.34) | n.a.§ |
| R split †† (%) | 62.3 (538) | 22.5 (424) | n.a.§ |
| CC1/2 | 0.70 (0.03) | 0.97 (0.16) | n.a.§ |
| CC* | 0.90 (0.25) | 0.99 (0.53) | |
| Anomalous CC‡‡ | n.a. | 0.06 (0.03) | n.a.§ |
| Completeness (%) | 98.1 (85.5) | 99.91 (100) | n.a.§ |
| Molecular replacement | |||
| Search-model PDB code§§ | 3n8g | 3n5k | |
| Rotation-function Z-score (ensemble 1/2) | 8.4 | 2.3/1.8 | |
| Translation-function Z-score (ensemble 1/2) | 10.6 | 11.5/20.4 | |
| Log-likelihood gain | 4295 | 991 | |
| Refinement | |||
| Resolution () | 602.80 (2.912.80) | ||
| No. of reflections | 41471 (3500) | ||
| Final R cryst/R free (%) | 30.4/34.3 (47.5/49.9) | ||
| No. of non-H atoms | |||
| Protein | 7671 | ||
| Ca2+ | 3 | ||
| K+ | 1 | ||
| AMPPCP | 31 | ||
| Total | 7706 | ||
| R.m.s. deviations | |||
| Bonds () | 0.006 | ||
| Angles () | 0.762 | ||
| Average B factors (2) | |||
| N domain | 97.7 | ||
| P domain | 97.6 | ||
| A domain | 114 | ||
| TM domain | 130.3 | ||
| Ca2+ | 113 | ||
| K+ | 91.0 | ||
| AMPPCP | 88.6 | ||
| Ramachandran plot | |||
| Most favoured (%) | 95.1 | ||
| Allowed (%) | 4.4 | ||
Numbers in parentheses are the percentage of the total collected frames.
Numbers in parentheses are the percentage of indexed frames.
The Zn2+-ATPase data set was not merged because the number of indexed patterns was too low to obtain meaningful merged data.
The according numbers of unique reflections when merged with Friedel pairs treated as separate reflections is 31576 (3262).
R
split is defined in CrystFEL as the R factor between data sets calculated from two randomly chosen halves of the data, corrected for the decrease in multiplicity caused by dividing the data into halves: R
split = 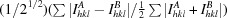 .
.
Determined from processing the data set with separated Friedel mates.
See 2 for details of how the search models were prepared to remove model bias.
