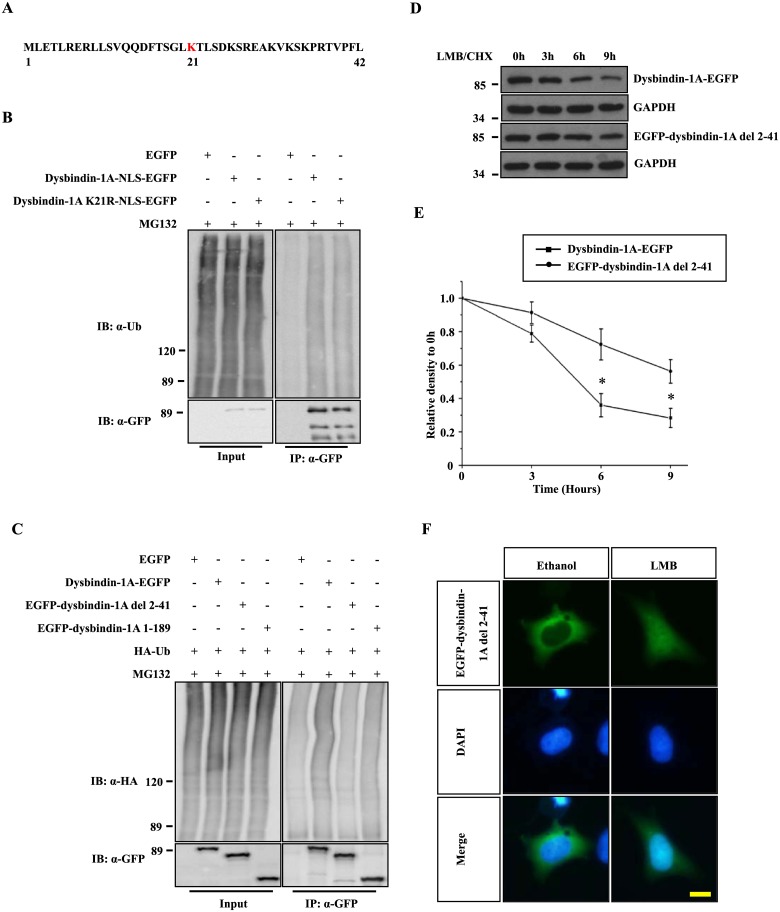
Fig 3. The N-terminal 2–41 amino acids of dysbindin-1A are important for its nuclear degradation.
(A) The N-terminal sequence of dysbindin-1A (amino acids 1–42). The sequence was scored using Ubpred and BDM-PUB, and potential ubiquitination sites are colored red. (B) The potential dysbindin-1A ubiquitination site, lysine 21, was mutated to arginine. The ubiquitination of K21R and WT forms of dysbindin-1A was examined. (C) Dysbindin-1A-EGFP, EGFP-dysbindin (residues 1–189) or EGFP-dysbindin-1A residue 2–41 deletion mutant were co-transfected with HA-Ub into HEK293 cells, respectively. After culturing for 24 hours, 10 μM MG132 was added, and the cells were then cultured for an additional 12 hours. After lysis, the proteins were immunoprecipitated using an anti-GFP antibody and immunoblotted with an HA antibody. (D) The EGFP-dysbindin-1A residue 2–41 deletion mutant was more stable in the nucleus. HEK293 cells expressing dysbindin-1A-EGFP or EGFP-dysbindin-1A residue 2–41 deletion mutant were treated with leptomycin B (20 ng/ml) for 1 hour. The cells were then treated with CHX for the indicated time. Finally, cell extracts were subjected to immunoblot analysis. (E) The band intensity of the dysbindin-1A-EGFP and EGFP-dysbindin-1A residue 2–41 deletion mutant is shown, relative to GAPDH. The values shown represent the means ± S.E. of three independent experiments. *, P<0.05; one-way ANOVA. (F) The localization of EGFP-dysbindin-1A del 2–41 under treatment of Ethanol or LMB. The nuclei were stained with DAPI; The bar represents 10 μm.