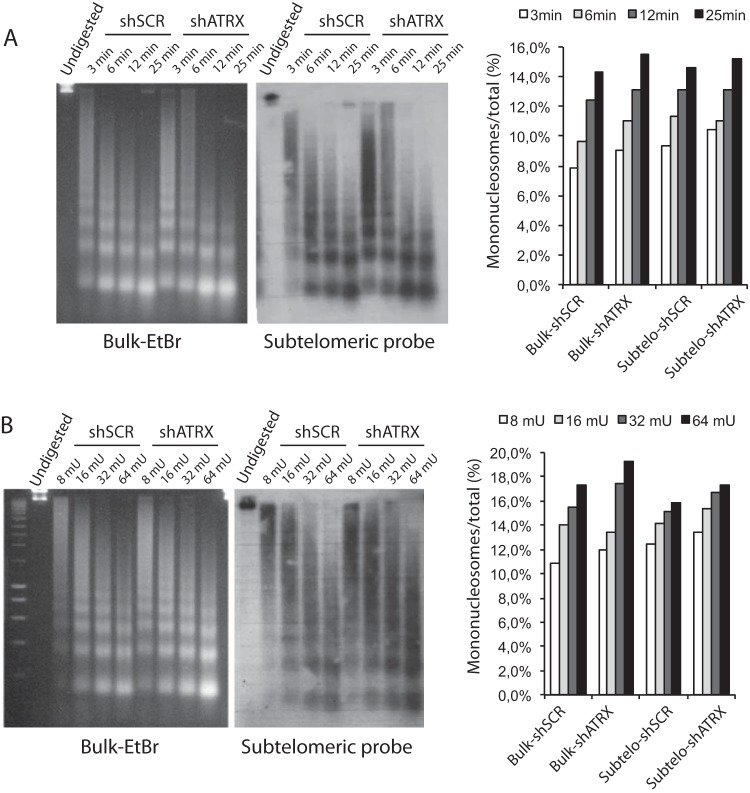
FIG 4.
shRNA-mediated inactivation of ATRX does not alter subtelomeric chromatin accessibility to MNase. (A) Chromatin isolated from 8-MG-BA glioma cells in which ATRX had been inactivated (shATRX) or not (shscrambled [shSCR]) was digested with MNase for the indicated times. (Left gel) Ethidium bromide (EtBr) staining of bulk chromatin. (Right gel) Southern blot with subtelomeric probe. (Far right) Quantification of the data. The signals obtained for mononucleosomes were normalized to the total signals measured for each time point (EtBr or Southern blot). (B) Chromatin samples from shATRX or shSCR 8-MG-BA cells were digested for 5 min with the indicated amounts of MNase (milliunits per microgram of DNA). (Far right) Quantification of the data.