Fig. 2.
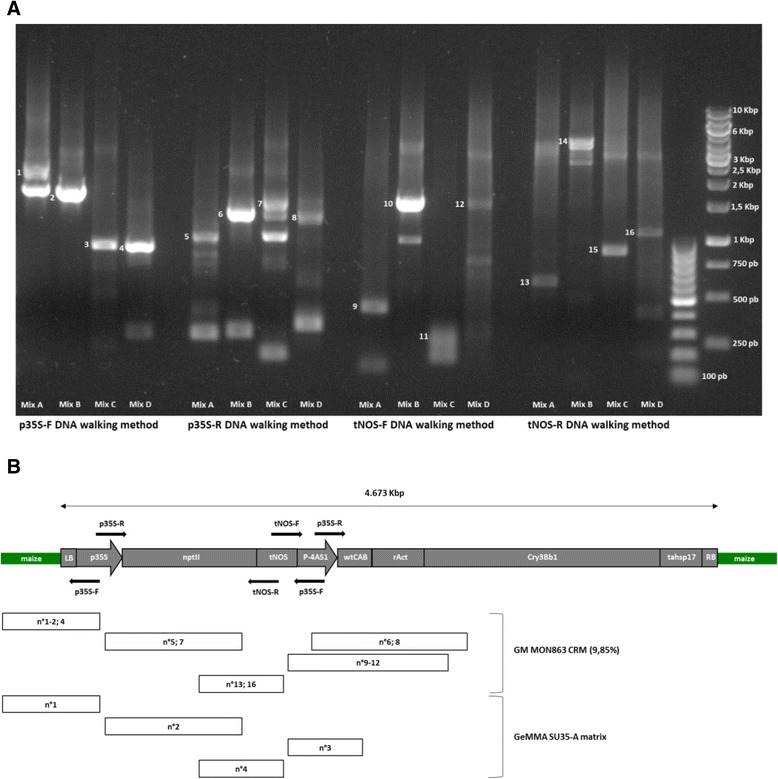
Application of the bidirectional p35S and tNOS DNA walking methods on GM maize matrices. a Visualisation of the obtained amplicons using the p35S and tNOS DNA walking methods applied on 100 ng of the GM MON863 maize CRM (9.85 %). For each method, four different DRT primer mixes (A-D) have been used. The analyzed amplicons are indicated by a numerotation going from 1 to 16. b For each DNA walking method, a schematic representation of the potential start position and direction, applied on the transgenic cassette of the GM maize MON863, is illustatred by the black arrows. Below the transgenic cassette, the sequence covering of the selected amplicons from the GM MON863 maize CRM (9.85 %) and the GeMMA proficiency test food matrix (GeMMA SU35-A) is schematically represented by rectangles. The corresponding amplicon numbering is indicated in the Fig. 2a and Additional file 6. LB (left border); p35S (CaMV 35S promoter); nptII (neomycin phosphotransferase II gene); tNOS (Agrobacterium tumefaciens nopaline synthase terminator); p4-AS1 (modified CaMV 35S promoter); wtCAB (Wheat major chlorophyll a/b binding protein gene); rAct (Rice Actin intron); Cry3Bb1 (synthetic Cry3Bb1 gene); tahsp17 (Wheat heat shock protein terminator); RB (right border); maize (maize genome) [Schema adapted from 27 and 30]
