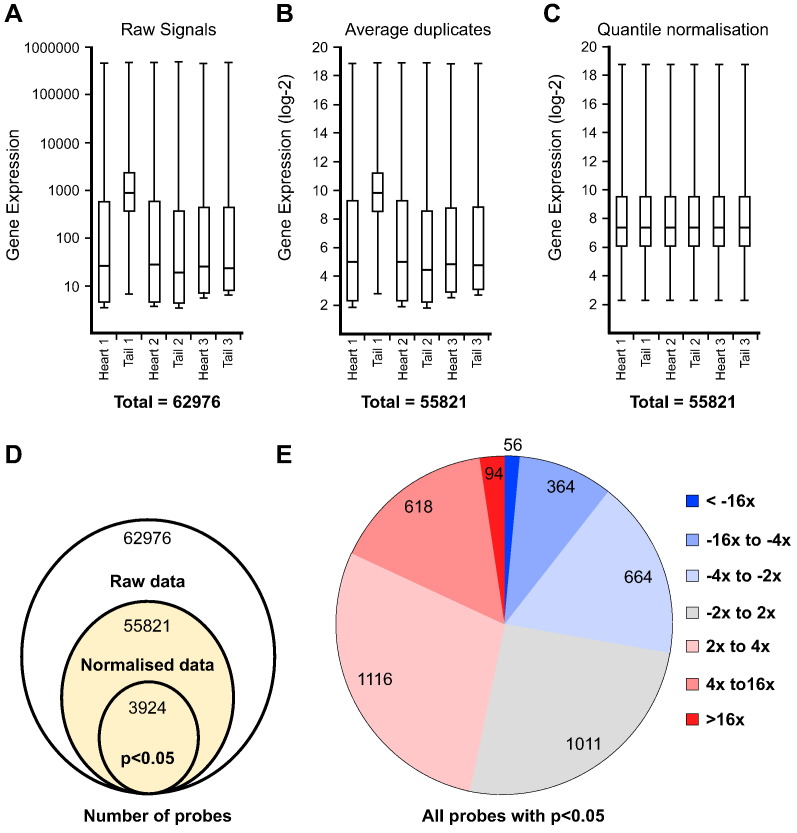
Fig. 2.
Normalisation and transcriptome-wide comparison between cardiac fibroblasts and tail fibroblasts. (A –C) Box-plots of the gene expression data at the end of each pre-processing and normalisation processing stage: (A) Single-channel signals extracted from Agilent Feature Extraction Software; (B) data pre-processing, duplicates averaging and log-2 transformation; and (C) quantile normalisation. After normalisation, all samples have identical distributions. (D) Venn diagram illustrating the initial number of entities in the raw data (62,976 genes), the reduction of data points through processing and normalisation (55,821 genes), and finally the pool of differentially-expressed entities with p-value < 0.05 (3924 genes). (E) Distribution of the level of gene expression fold-changes within the pool of 3924 differentially-expressed entities.