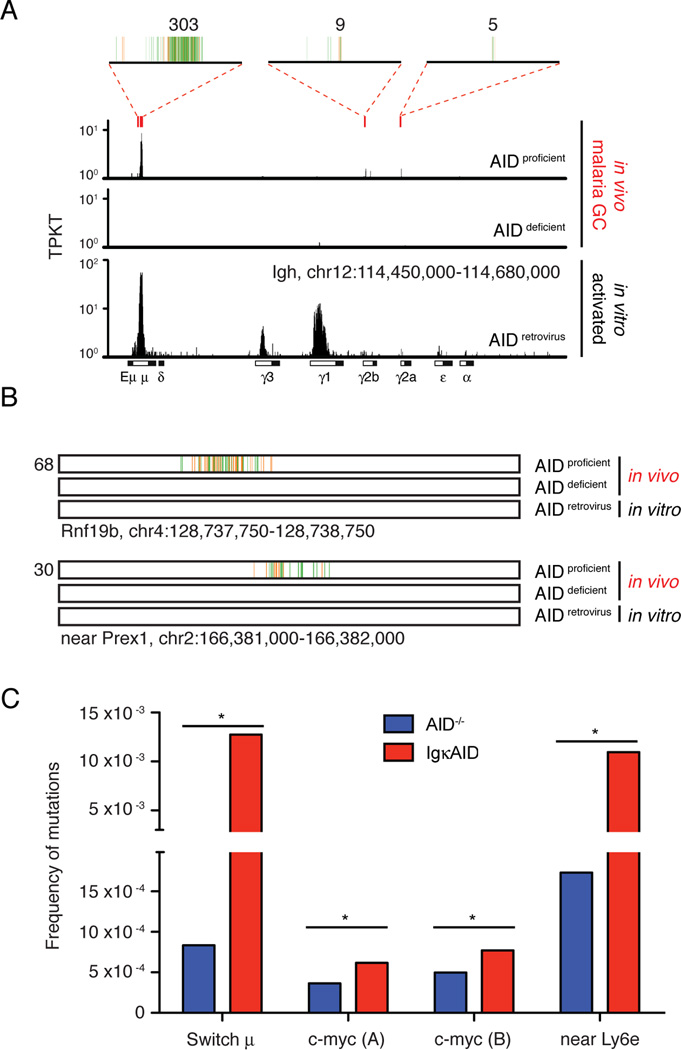
Figure 4. AID contributes to Plasmodium induced DNA damage.
A- Translocations at the physiologic AID target Igh. Red rectangles indicate the location of AID hotspots, and 4kb regions at these hotspots are magnified on top, where vertical lines each represent a unique translocation event. Numbers on top indicate the total of translocations at each hotspot region. Rearrangements obtained in cultured B cells retrovirally expressing AID are shown for comparison (Klein et al., 2011). TPKT is the normalized number of translocations per kilobase per thousand translocations in the library.
B- Examples of non-Igh hotspots of AID-dependent translocation induced by malaria. One kb region is shown, with each vertical line representing a unique translocation. Numbers on the left indicate the total of translocations.
C- Mutational analysis of malaria GC B cells DNA by MutPE-seq. For c-myc, two adjacent regions in intron 1 were analyzed. P value is *<0.000001 for all (one-tailed Student’s T-test). See also Table S1 and Figure S4.