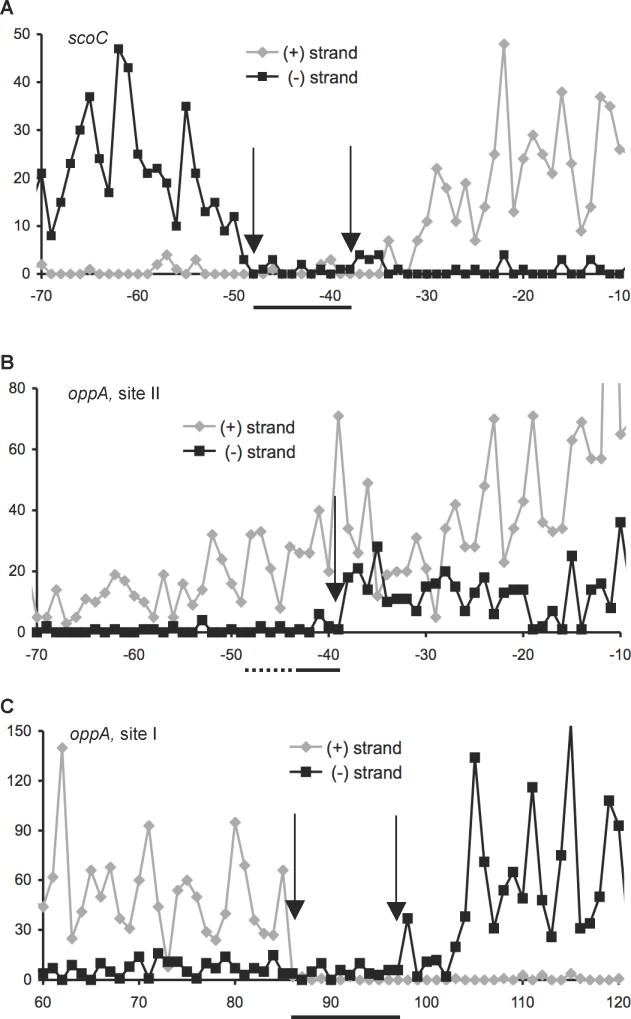
Fig. 2.
Coverage maps of CodY-binding sites in the (A) scoC and (B and C) oppA regulatory regions using strand-specific counting of 5’ nucleotides of Illumina sequencing reads obtained in IDAP-Seq experiments. The coverage of the (+) and (−) strands is shown in grey and black, respectively. Arrows indicate boundaries of coverage gaps corresponding to the boundaries of core binding sites at the CodY concentration used. The core site location is also indicated by horizontal lines below x-axes; the 5’ end of the oppA site II was not determined. Forty nM CodY was used for affinity purification of DNA fragments in presented experiments; the boundaries of the core sites, mentioned in the text, were deduced from multiple experiments using a range of CodY concentrations. Non-uniform coverage is likely due to varying position-specific efficiency of DNA shearing during sonication and to non-uniform size of DNA fragments. Coordinates are reported with respect to the transcription start points. The results shown here were deduced from data provided as supplementary material in a previous publication (Belitsky and Sonenshein, 2013) but were not previously analyzed.