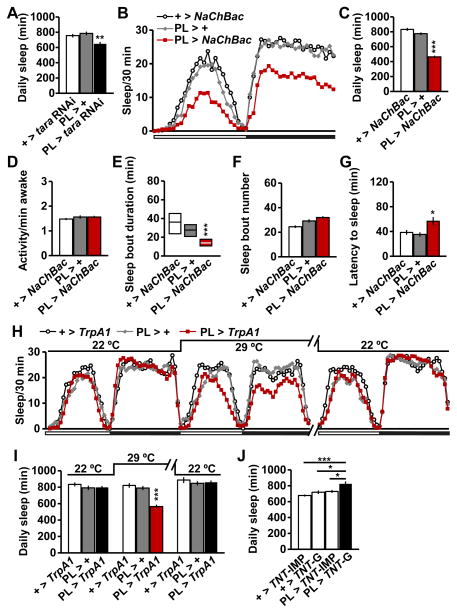
Figure 5. TARA Regulates Sleep in CycA-Expressing PL Neurons that Define a New Arousal Center.
(A) Total daily sleep of female flies of the indicated genotypes (n=32). (B) Sleep profile for female flies expressing the NaChBac sodium channel under the control of PL-Gal4 (PL > NaChBac) and parental controls (n=48–63). (C–G) Total daily sleep (C), waking activity (D), sleep bout duration (E), sleep bout number (F), and latency to sleep after lights off (G) for the flies shown in (B). (H) Sleep profile for female flies expressing TrpA1 under the control of PL-Gal4 (PL > TrpA1) and parental controls (n=16–32). Flies were monitored at 29°C, which activates the TrpA1 channel, and at 22°C, which inactivates the TrpA1 channel. Sleep profile for the second day at 29C is omitted for simplicity. (I) Total daily sleep for flies shown in (H). (J) Female flies expressing functional tetanus toxin under the control of PL-Gal4 (PL > TNT-G) exhibited a significant increase in sleep relative to flies expressing inactive tetanus toxin (PL > TNT-IMP) or those carrying either form of tetanus toxin transgene without the PL driver (+ > TNT-G and + > TNT-IMP) (n = 30–32). Mean ± SEM is shown. * p < 0.05, **p < 0.01, ***p < 0.001, ns: not significant, Dunnett post hoc test relative to parental controls (A, C–D, F–G, I) or PL > TNT-G flies (J); Kruskal-Wallis test (E). See also Figure S5.