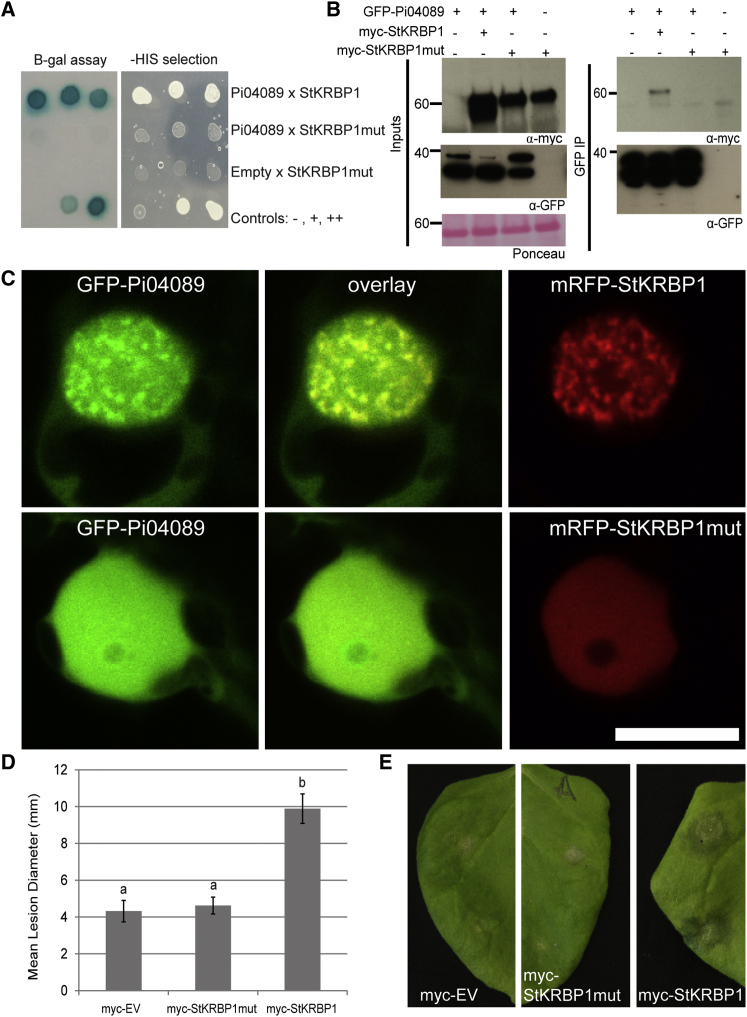
Figure 7.
A KRBP1 Mutant No Longer Interacts with Pi04089 and Fails to Promote P. infestans Colonization.
(A) Yeasts co-expressing the StKRBP1 with Pi04089 grow on -histidine (-HIS) medium and have β-galactosidase (B-gal) activity, whereas yeasts co-expressing StKRBP1 mutant (StKRBP1mut) do not. The control combination of StKRBP1mut with the empty prey vector indicates that it does not autoactivate. Controls show no (-), weak (+), and strong (++) interactions.
(B) Co-immunoprecipitation confirmed that Pi04089 specifically interacted with StKRBP1 and not with the mutated protein.
(C) Single optical sections of nuclei of cells co-expressing GFP-Pi04089 with either the wild-type StKRBP1 or the mutated form, showing that GFP-Pi04089 is only removed from the nucleolar periphery and relocated to speckles by the wild-type, and also that the mutated StKRBP1 does not locate to nuclear speckles. Scale bar represents 10 μm.
(D) Graph shows that the mean diameter of P. infestans lesions following Agrobacterium-mediated expression of myc-StKRBP1 mutant was significantly decreased compared with the expression of the wild-type myc-StKRBP1 and did not significantly differ from the empty vector (myc-EV) control (p < 0.001, ANOVA, as indicated by lowercase letters). Error bars are SE, and the graph represents the combined data from three biological reps (n = 72 per construct).
(E) Example infection sites on N. benthamiana leaves.