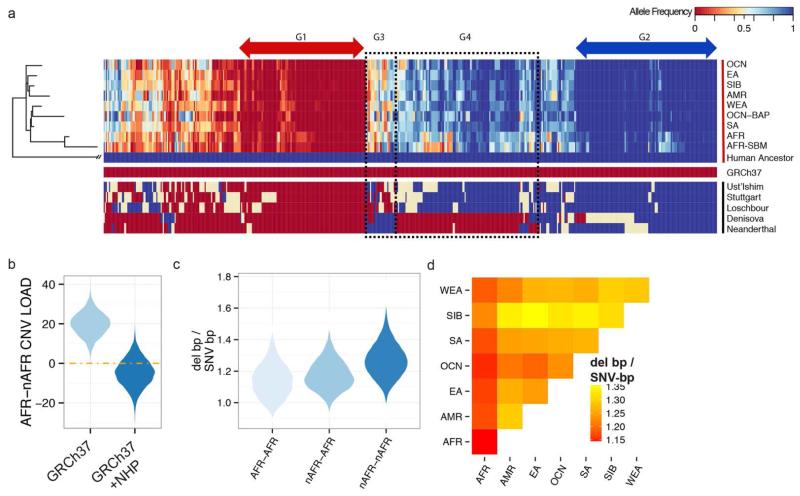
Fig. 5. The ancestral human genome and CNV burden.
(A) A heatmap of the allele frequency of 571 (1.55 Mbp) nonrepetitive sequences absent from the human reference genome yet segregating in at least one human population ordered in humans by a maximum likelihood tree (49). Four groups of interest are highlighted: G1 – ancestral sequences that have almost been completely lost from the human lineage, G2 – ancestral sequences that are largely fixed but rarely deleted (also absent in human reference), G3 – ancestral sequences that have become copy number variable since the divergence of humans and Neanderthals/Denisovans ~700 kya, and G4 – sequences potentially lost in Neanderthals and Denisovans since their divergence from humans. (B) The resulting distributions of 10,000 block-bootstrapped estimates of the difference in load between African (AFR) and non-African (nAFR) populations considering only the reference genome (GRCh37) and supplemented by sequence absent from the human reference genome (GRCh37 + NHP) included (see text for details). (C) Violin plots of the distribution of the ratio of deletion base pairs to SNV base pairs differing between every pair of African individuals (AFR-AFR), all pairs of non-African individuals (nAFR-nAFR) and every non-African, African pair (nAFR-AFR). (D) Heatmap representation of the mean ratio of deletion to SNV base pairs differing between individuals from pairs of populations.