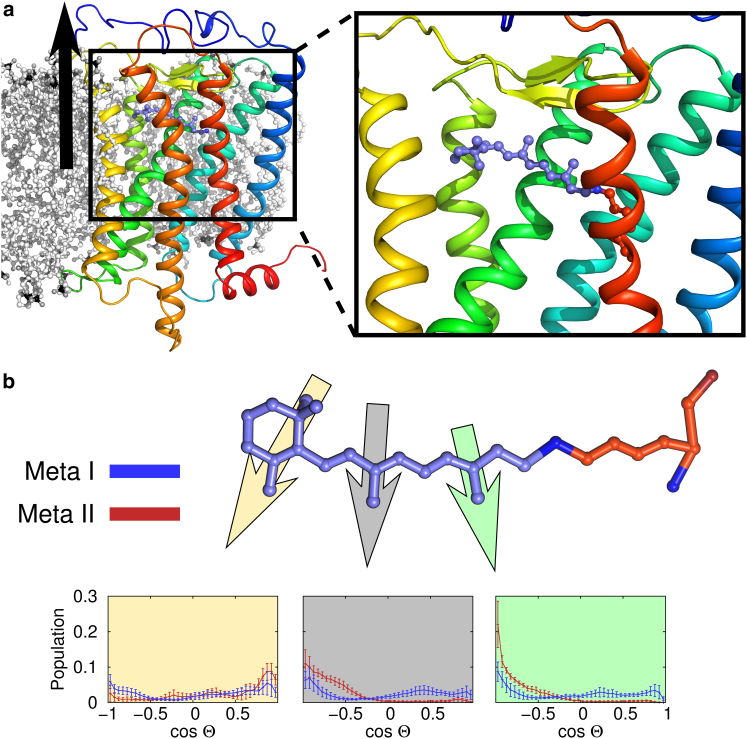
Figure 2.
Retinal dynamics distinguish the Meta-I and Meta-II ensembles. (a) Illustration showing rhodopsin in a lipid bilayer. The protein is shown as a rainbow colored cartoon with the N-terminus colored blue. Lipids are shown in a black/white ball-and-stick representation (note several lipids are removed for clarity). The black arrow denotes the membrane normal. The inset shows a close-up view of retinal (purple ball-and-sticks) in the binding pocket. It is covalently attached to Lys2967.43 (orange) by a Schiff-base linked nitrogen. (b) Retinal orientations. A morphed retinal is shown colored as in (a). Its orientation was monitored using the three vectors illustrated by yellow, gray, and green arrows. Specifically, we computed the dot-product between a vector drawn from either: the C1–C5 atoms (yellow), the C9–C19 atoms (gray), or the C13–C20 atoms (green) and the membrane normal (black arrow in a). This angle was histogrammed over the entire Meta-I (blue data) or Meta-II (red data) ensemble of trajectories. The average histograms are shown below, with error bars representing the standard error. To see this figure in color, go online.