Figure 3.
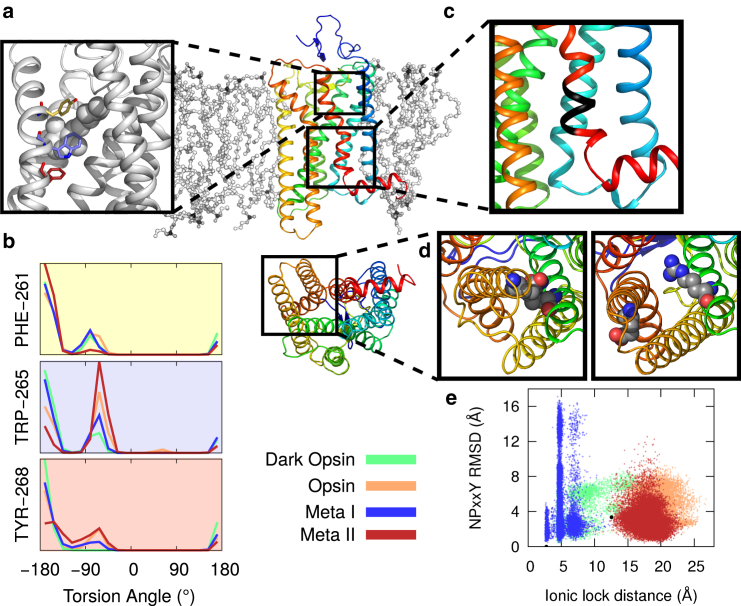
Dynamics of biologically distinguished structural motifs. (Center) Rhodopsin is shown as a rainbow cartoon embedded in a lipid bilayer. The bottom view shows the protein’s cytoplasmic face. (a) Illustration showing retinal (white spheres) in its binding pocket and rotamer toggle residues: Phe2616.44 (pale yellow), Trp2656.48 (pale violet), and Tyr2686.51 (dark red). (b) Histograms of the population of torsion angles for each of the three toggle switch residues. Data is colored by the simulation ensemble. (Note: a version of this figure with error bars is depicted in Fig. S8.) (c) Close-up view of the NPxxY motif. (d) The ionic lock. Arg1353.50 and Glu2476.30 are shown forming a salt bridge (left) as in the inactive crystal structure (PDB ID: 1U19) or in an open conformation (right) as in the active crystal structure (PDB ID: 3PXO). (e) NPxxY motif versus ionic lock dynamics. The NPxxY RMSD from the inactive crystal structure (y axis) is shown versus the ionic lock distance (x axis). Every frame (sampled at a rate of 1 ns) is shown colored according to ensemble. To see this figure in color, go online.