Fig. 4.
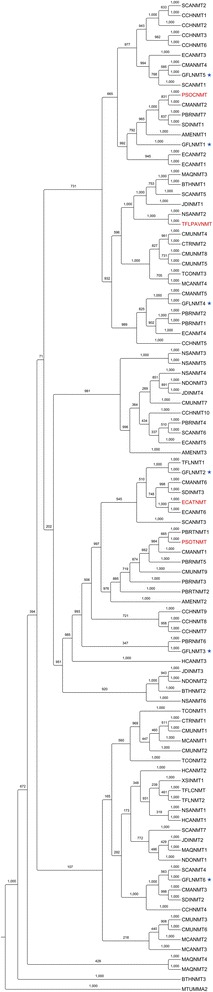
Phylogenetic analysis of N-methyltransferase (NMT) gene candidates from twenty BIA-accumulating plant species. Red text denotes characterized genes or enzymes used as tBLASTn queries for transcriptome mining. Black text denotes uncharacterized gene candidates identified through mining (>40 % identity to queries). Bootstrap values for each clade were based on 1000 iterations. Each candidate is labeled with respective species abbreviation (e.g. AME, Argemone mexicana; see Table 1) and candidate number (e.g. NMT1). Each query is labeled according to species (additional species: PSO, Papaver somniferum) and specific NMT function (CNMT, coclaurine N-methyltransferase; PAVNMT, pavine N-methyltransferase; TNMT, tetrahydroprotoberberine N-methyltransferase; see Fig. 1). Outgroup is mycolic acid synthase from Mycobacterium tuberculosis (MTUMMA2). NMT candidates from Glaucium flavum tested for catalytic activity are indicated with asterisks. Amino acid sequences for candidates, queries, and outgroups are found in Additional file 6
