Fig. 2.
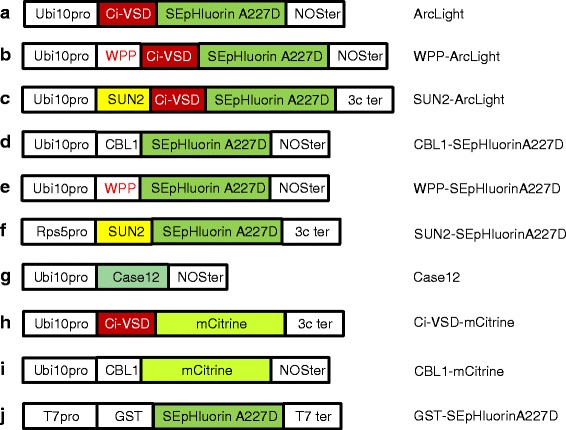
Constructs used in this study. The construct letters (a-f) correspond to the diagram letters in Fig. 1. SEpHluorinA227D, Ci-VSD and Case12 are defined in the text. The CBL1 motif is a 12 amino acid sequence from the CBL1 protein that contains a myristolated glycine and a palmitolated cysteine, which tether the fluorescent fusion protein to the cytoplasmic surface of the plasma membrane [25]. The WPP sequence, which contains a Trp(W)-Pro(P)-Pro motif that is highly conserved in all land plants [22], consists of amino acids 28–131 of Arabidopsis RANGAP1 and is sufficient for targeting fusion proteins to the outer nuclear membrane [23]. The SUN2 protein, which is 455 amino acids in length, has one transmembrane domain that can localize SUN2-fusion proteins at the inner nuclear membrane surface [44, 45] . In constructs (a-f) and (g-i), the gene encoding the fluorescence reporter is under the control of the ubiquitously-expressed Ubi10 plant promoter [39]. Construct F contains the root-specific Rps5 promoter [40]. Ci-VSD-mCitrine corresponds to VSFP3.1_mCitrine [28]. The constructs (a-i) contain either the nopaline synthase (NOS) or 3C transcriptional terminator. Construct (j) is designed for expression of GST-tagged SEpHluorinA227D in E. coli and contains the phage T7 promoter and terminator. The amino acid sequences of SEpHluorinA227D and environmentally-insensitive monomeric (m)Citrine compared to wild-type GFP and SEpHluorin are shown in Additional file 1: Figure S1. The constructs are not drawn to scale
