Fig. 2.
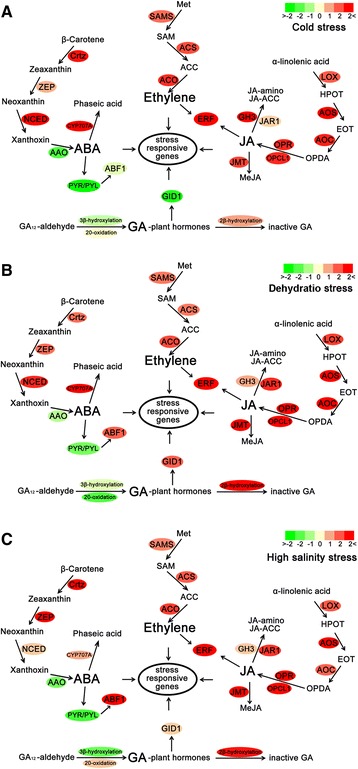
Expression and pathway distribution of phytohormone metabolism and signal transduction-related genes in M. falcata. a Expression pattern of key genes under cold stress. b Expression pattern of key genes under dehydration stress. c Expression pattern of key genes under high salinity stress. Additional abbreviations are as follows: Crtz (beta-carotene 3-hydroxylase); ZEP (zeaxanthin epoxidase); NCED (9-cis-epoxycarotenoid dioxygenase); AAO (abscisic-aldehyde oxidase); PYR/PYL (PYRABACTIN RESISTANCE1 / PYRABACTIN RESISTANCE 1-LIKE); ABF1 (ABRE-binding factors 1); GID1 (GA-INSENSITIVE DWARF1); SAMS (S-adenosylmethionine synthetase); ACS (1-aminocyclopropane-1-carboxylic acidsynthetase); ACO (1-aminocyclopropane-1-carboxylic acid oxidase); ERF (ethylene response factor); LOX (lipoxygenase); AOS (allene oxide synthase); AOC (allene oxide cyclase); OPR (12-oxo-phytodienoic acid reductase); OPCL1 (OPC-8:0 CoA ligase 1); JMT (jasmonate O-methyltransferase); and JAR1 (Jasmonate response 1). Filled colours correspond with their degree of regulation by the stresses
