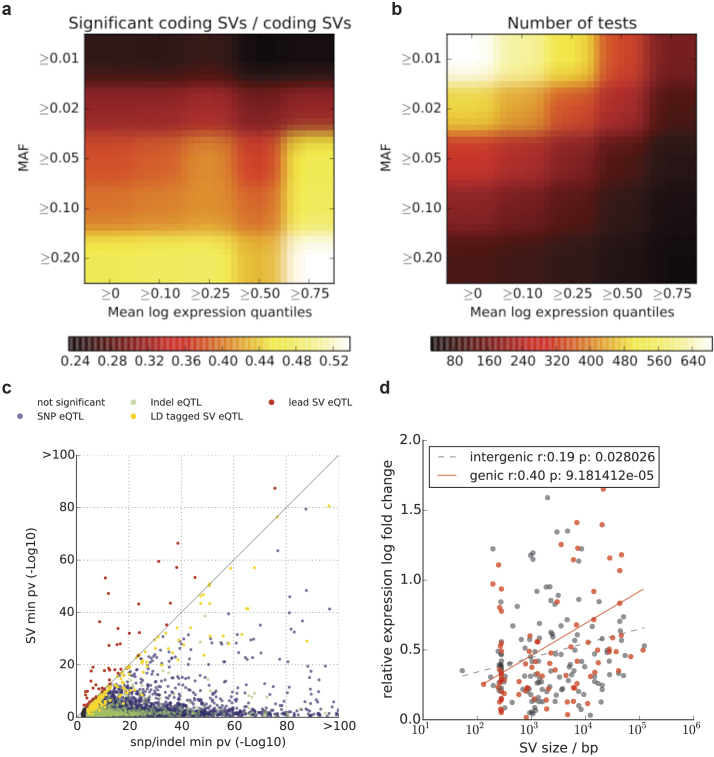
Extended Data Figure 8. SV-eQTL analysis.
a, SV-centric eQTL analysis of coding SVs. Shown is the proportion of coding SVs that are eQTLs as a function of the minimum VAF and the expression quartile. b, Total number of coding SVs for corresponding filters. Common SVs (VAF > 0.2) in highly expressed genes (>75% quantile) are very likely to correspond to SV-eQTLs (54%, see also Supplementary Table 8). c, For all genes with significant eQTLs (FDR < 10%), shown are raw P-values considering only SNPs (x axis) or only SVs (y axis). Genes with (strict lead) SV-eQTLs are shown in red. Genes with a SNP lead eQTL that is in linkage with an SV (r2 > 0.5) are shown in orange. SNP lead eQTLs without an SV in LD are shown in blue. d, Relative eQTL effect sizes for genetic and intergenic SV eQTLs (n = 239) either with an SV-eQTL or an LD tagged SV (in log abundance scale). Shown are regression trends for both genic and intergenic SV eQTls. For genetic eQTLs, a clear relationship between SV effect size is found. For example, genic SVs >10 kb have threefold larger effect sizes compared to genic SVs < 1 kb; P = 0.004; t-test.