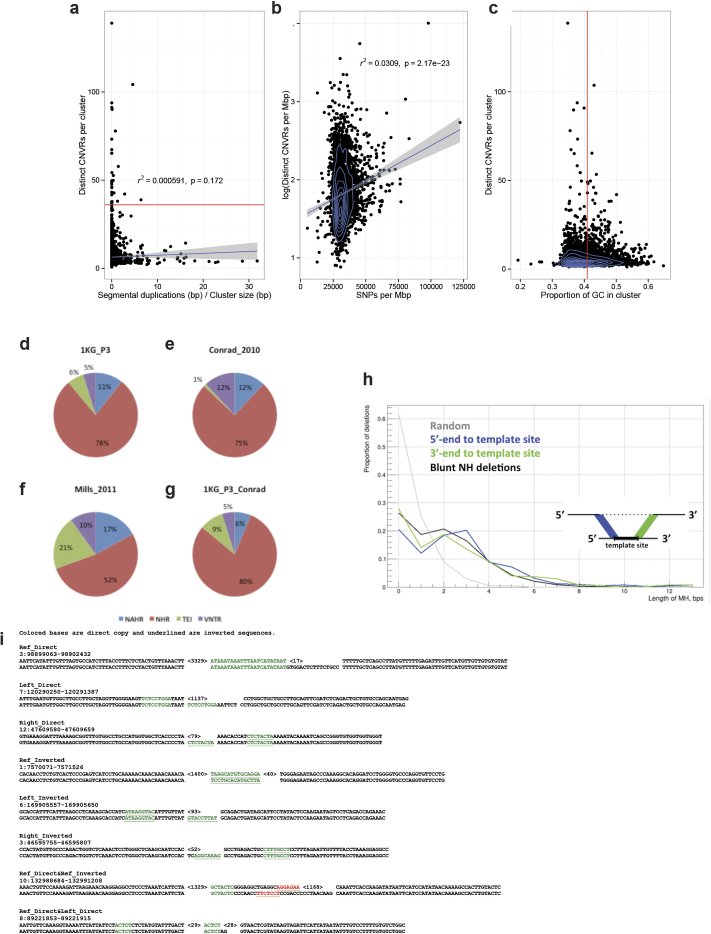
Extended Data Figure 9. SV clustering and breakpoint analysis.
a–c, Extensive clustering of recurrent SVs into CNVRs appears unrelated to the extent of segmental duplications (a) and is only partially correlating with SNP diversity (b) and GC content (c). Breakdown of SV mechanism classifications based on criteria from two earlier studies (refs 6, 40). Shown are results for deletions with nucleotide resolved breakpoints. BreakSeq was used for mechanism inference. d, 1KG_P3: breakdown for our 1000 Genomes Project phase 3 SV callset using classification criteria from ref. 6. e, Conrad_2010: summary of mechanism classification results published in ref. 40. f, Mills_2011: summary of mechanism classification results published in ref. 6. g, 1KG_P3_Conrad: Breakdown for our 1000 Genomes Project phase 3 SV callset using classification criteria from ref. 40. Mechanism classification was pursued using four different categories. Blue, non-allelic homologous recombination (NAHR); green, mobile elements inserted into the reference genomes (appearing deleted in this analysis); red, non-homology-based rearrangement mechanisms (NHR), such as NHEJ, microhomology-mediated end-joining and microhomology-mediated break-induced replication (involving blunt-ended deletion breakpoints or breakpoints with microhomoloy); purple, expansion or shrinkage of variable numbers of tandem repeats (VNTRs). TEI, transposable element insertion (equivalent with MEI). h, Distribution of lengths of micro-homology (MH) for complex SVs, measured between deletion and corresponding template sites boundaries. Simple deletions, which based on BreakSeq were inferred to be formed by a non-homology-based SV formation mechanism, such as NHEJ and microhomology-mediated break-induced replication (Supplementary Table 3), are shown as an additional control (here denoted ‘blunt NH deletions’). i, Origins of inserted sequences in complex deletions inferred by split read analysis. This figure depicts examples for each class shown in Supplementary Table 13.