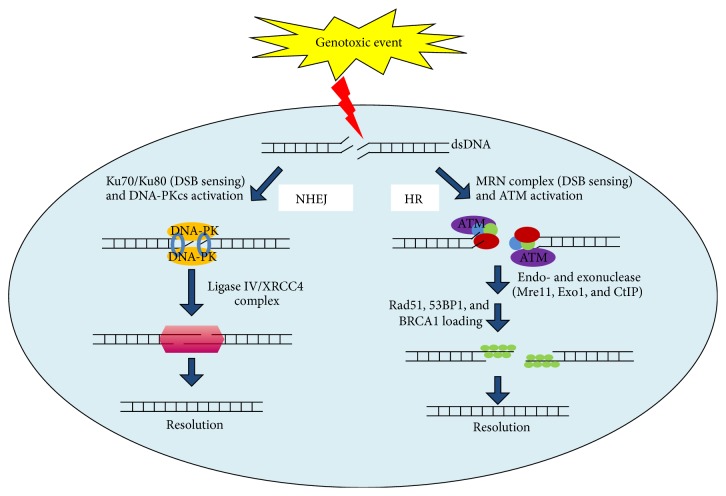
Figure 1.
DSBs repair mechanisms. On the left part NHEJ. DSBs are identified by the ring-shape heterodimer Ku70/Ku80 which binds DNA broken ends and recruits the DNA-PKcs (DNA-dependent protein kinase catalytic subunit). This complex stabilizes DNA ends allowing a ligation carried out by XRCC4 and Ligase IV complex that finally reattaches the broken DNA. On the right part HR. The ATM kinase is recruited to DSB via an interaction with the MRN (Mre11-Rad50-Nbs1) complex, RNF8 and RNF168. Once ATM becomes activated, it phosphorylates multiple substrates including endo- and exonuclease (such as Mre11, Exo1, and CtIP) that are coated with ssDNA. Moreover ssDNA regions attract also Rad51 and other associated proteins (53BP1, BRCA1, etc.) which collectively assure new DNA synthesis. Defects of these mechanisms/cooperation lead to genomic instability, which in turn mediates tumor cell growth and progression.