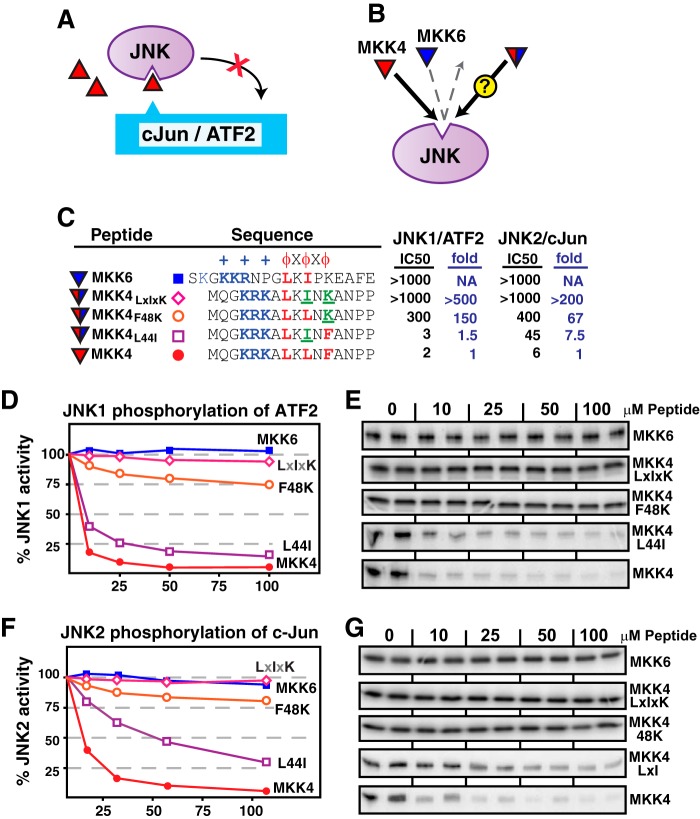
FIGURE 2.
Effect of “MKK6-like” hydrophobic residue substitutions on MKK4-JNK binding. A, peptide competition assay. D-site peptides (triangles) were used to inhibit JNK phosphorylation of the D-site-containing substrates c-Jun or ATF2. B, the MKK4 D-site peptide (red) binds with high affinity to JNK and inhibits JNK-mediated phosphorylation of D-site-containing substrates. The MKK6 D-site peptide (blue) does not bind to JNK and therefore does not inhibit JNK-mediated phosphorylation. The experiment was designed to measure the binding of MKK4 peptides that were substituted with residues taken from MKK6. Will these hybrid D-site peptides bind to JNK or not? C, sequence of D-site peptides used in this experiment; all sequences used in this paper are human. Substituted residues are indicated in green and underlined. To the left of the sequences, the IC50 (the peptide concentration that inhibits phosphorylation by 50%; see “Experimental Procedures” for more details) for each kinase/substrate combination tested is shown. For the mutant MKK4 peptides, the -fold increase in IC50, normalized to wild-type MKK4, is also shown. Lower IC50 values indicate stronger D-site·JNK binding; thus, “-fold increase in IC50” quantifies the factor by which JNK binding has been reduced by the mutations indicated. D, purified GST-ATF (1 μm) was incubated with purified active JNK1 (∼50 nm) and [γ-32P]ATP for 20 min in the absence or presence of the specific concentrations of the indicated peptides. In the graph, results are plotted as percent phosphorylation relative to that observed in the absence of any added peptide. Phosphate incorporation into ATF2 was analyzed by SDS-PAGE and quantified on a PhosphorImager. Data are the average of 3–8 experiments, with duplicate data points in each experiment. The S.E. between experiments was typically <10% of the mean; error bars are omitted for visual clarity. E, representative autoradiogram of experiments averaged and graphed in D. F and G, similar to D and E, except the kinase was JNK2, and the substrate was GST-c-Jun.