Fig. 6.
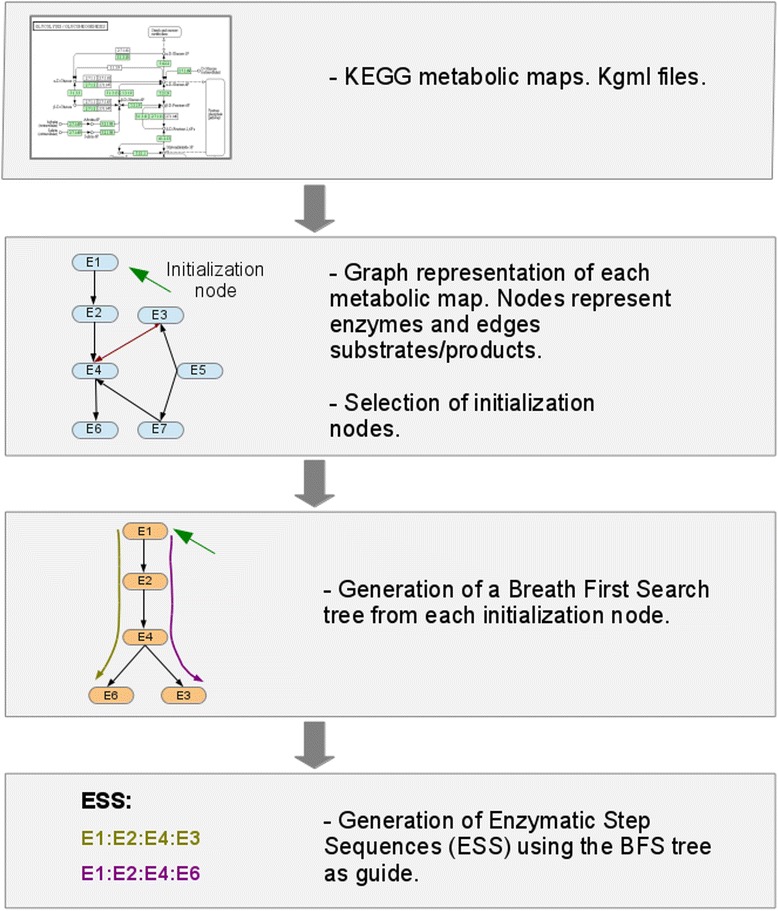
Strategy to generate the Enzymatic Step Sequences (ESS) from KEGG kgml files using the Breadth First Search (BFS) algorithm. First the kgml files are retrieved and directed graphs representations created. The nodes, represents genes or groups of genes (isozymes or complexes) that catalyze the reactions and the edges represent a compound that is a substrate from one reaction and a product for the next. Next a set of initialization nodes are selected using two criteria: a node which substrate is not catalyzed by another enzyme in the metabolic map; and, a node which substrate comes from another metabolic map and with two or less neighbors in the graph. Then, the initialization nodes are used as root for the construction of the BFS tree. Finally, the tree is used as guide to the construction of the nrESS. From each leaf (terminal node), the path is traced backwards until reach the root
