Fig. 6.
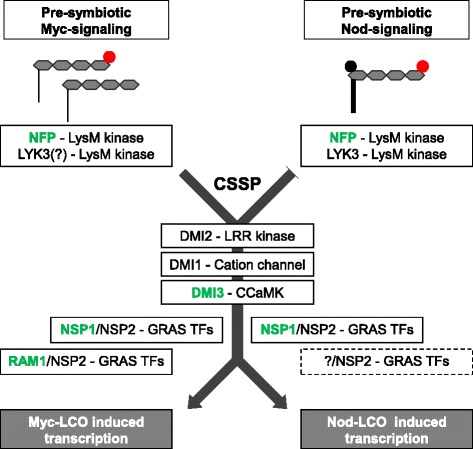
Model for the pre-symbiotic activation of host genes by microbial LCOs. This model integrates gene expression responses towards Nod- and Myc-LCOs in the wild type and in four symbiotic mutants at concentrations of 10-7/-8 M (Figs. 2, 4, 5). The two major pathways for a pre-symbiotic transduction of Nod- and Myc-LCO signals are shown. On top of the figure, LCO structures are depicted with four N-acetylamine residues. Sulfate groups of Nod- and sMyc-LCOs are shown in red, the acetate group typically present on Nod-LCOs in black, and differences in fatty acids by a different width. All components of signal perception and transduction studied in this work are shown in green. Other components of signal transduction reported in the literature are indicated in black. The partial dependency of Myc-LCO perception on LYK3 at high concentrations (10−5 M, [8]) is indicated by a question mark, since it is unknown if LYK3 is relevant at 10-7/-8 M as well. Whereas NSP1 and RAM1 are both essential for an up-regulation of Myc-LCO related genes not responding to Nod-LCOs, NSP1 is primarily required for the induction of genes by Nod-LCOs. The possible presence of additional GRAS TFs involved in activating Nod-LCO related gene expression is indicated by a dashed line. Abbreviations: CSSP, common symbiotic signaling pathway; CCaMK, Ca2+/calmodulin-dependent protein kinase; LRR, leucine-rich receptor; TF, transcription factor
