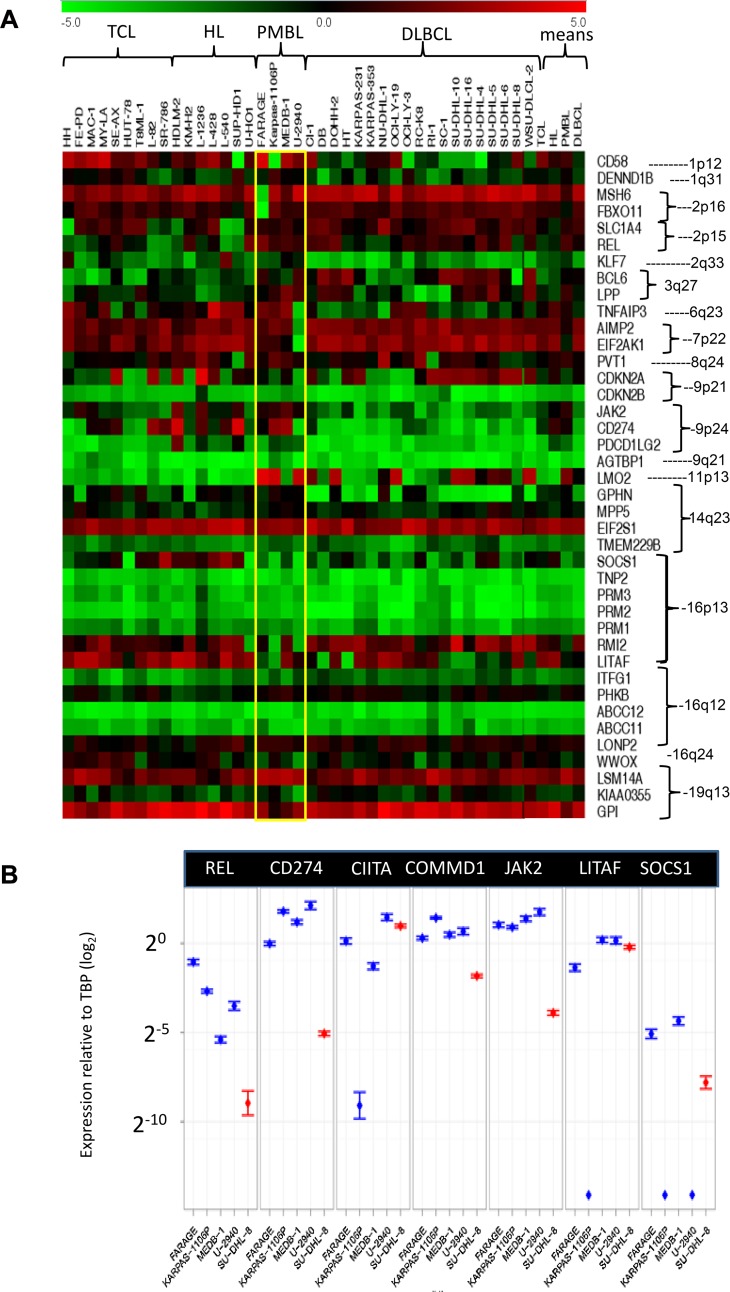
Fig 4. Gene expression at genomically rearranged loci.
A: Microarray expression data for genes implicated at CNV (mainly deletions). Note, for example gene silencing accompanying focal biallelic deletions at multiple loci: including, CD58 at 1p12 in KARPAS-1106P; MSH6 and FBXO11 at 2p26 in FARAGE; TNFAIP3, AIMP2 at 6q23 in U2940; EIF2AK1 at 7p22 in U-2940; CDKN2A at 9p21 in KARPAS-1106P and U-2940; CD274 at 9p24 in U-2940; SOCS1 at 16p13 in KARPAS-1106P and U-2940; and KIAA0355 at 19q13 in U-2940. B: Shows qPCR expression of select target genes at recurrent PMBL amplicons (2p15, 9p24) and a deletion (16p13) set against TBP reference in cell lines FARAGE, KARPAS-1106P, MEDB-1, and U-2940 (PMBL) shown blue, alongside reference SU-DHL-8 (DLBCL) shown red. Diamonds indicate undetectable expression. Quantitative data were verified by twofold or more biological replication.