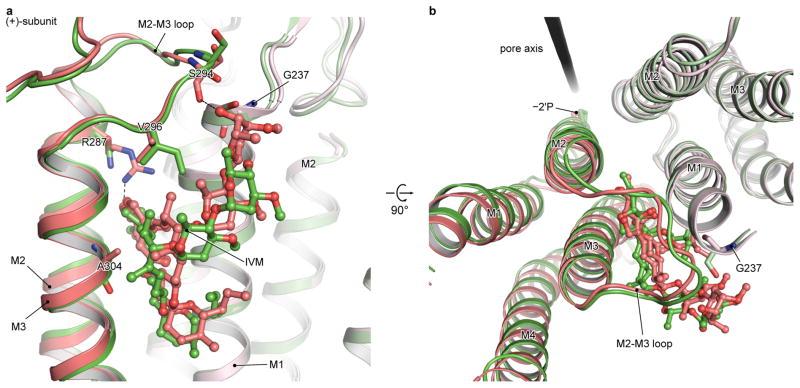
Extended Data Figure 6. Comparison of ivermectin binding site in GlyR (red) and GluCl (green), viewed in parallel to the membrane (a) or from the extracellular side (b).
The (+)subunits are shown in darker colors. The residue corresponding to Arg287, which forms a hydrogen bond with the ivmermectin in GlyR, is an asparagine (Asn264) in GluCl. The corresponding residue of Val296 in the M2-M3 loop of GlyR is an isoleucine (Ile273) in GluCl, whose larger side chain prevents the upper tip of ivermectin from approaching and interacting with the main chain oxygen atom of Ser721 in the M2-M3 loop (Ser294 in GlyR). The Gly237 in the M1 and Ala304 in the M3 of GlyR are Ser217 and Gly281 in GluCl, respectively. Such differences on side chains weaken or strengthen the interaction of ivermectin with M3 or M1 in GlyR, respectively, in comparison to that in GluCl.