Fig. 2.
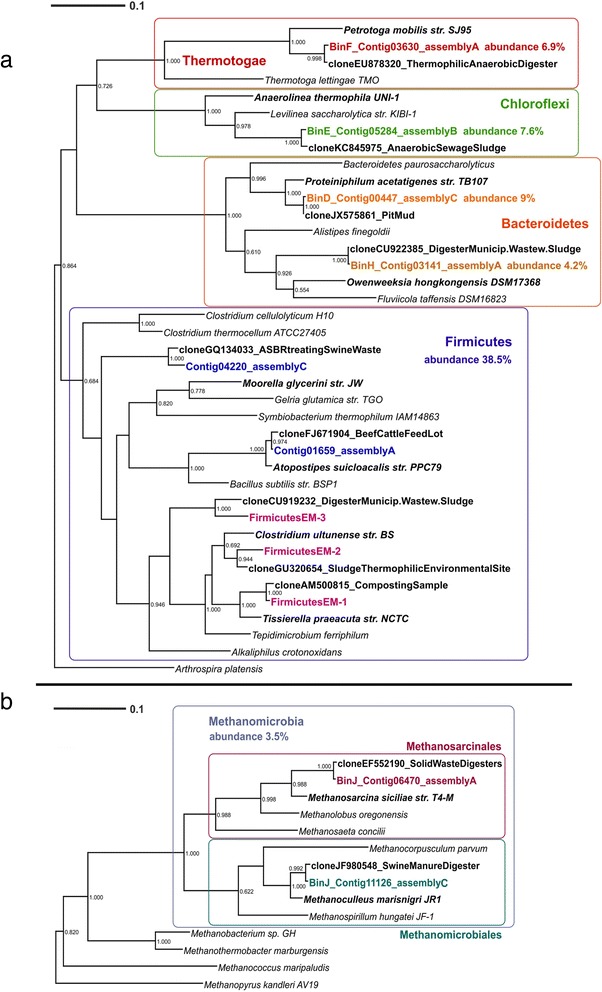
16S rDNA bacterial and archaeal phylogenetic trees. Phylogenetic trees of (a) Bacterial and (b) Archaeal 16S rDNA sequences. Although the binning was only performed with contigs of assembly A, most of the binned 16S rDNA sequences were also assembled in assemblies B and C. If so, for the tree the longest assembled sequence was chosen. 16S rDNA sequences not assembled in assembly A, but in B or C, or detected by EMIRGE are also included. Colored: sequences obtained from metagenomic reads. Assignment to metawatt bins is indicated if applicable. Reference sequences in bold: top hits in blast search against NCBI non-redundant nucleotide collection, bold + italics: top hits in blast search against NCBI reference RNA sequences. Additional reference sequences in the Bacteria tree represent genera detected in other anaerobic digesters. 16S rDNA sequences of Arthrospira platensis and Methanopyrus kandleri were chosen as outgroups, respectively. Bootstrap values at nodes are obtained from 500 replicates and are only shown for branches with at least 50 % support (values > 0.499). The scale bar represents 0.1 nucleotide substitutions per site. Accession numbers of reference sequences are available in Additional file 2: Table S7
