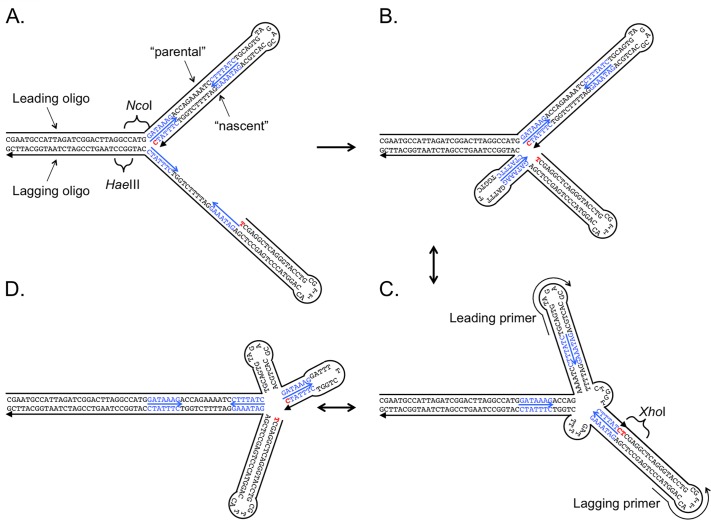
Fig 2. An in vitro construct to test for fork reversal and cross-strand ligation.
An artificial replication fork was generated using commercially synthesized oligonucleotides (sizes of 100 and 99 nucleotides, with a 5’-phosphate on the 99 nucleotide lagging oligonucleotide). (A) To facilitate assembly of the fork from its component strands, the “parental” and “nascent” strands of the leading arm of the duplex, and likewise, the two strands of the lagging arm of the replication fork were covalently joined by a short linker. Complementary sequences in the two, 30 base, single stranded tails allow the fork to assemble in vitro. Features engineered into the two oligos include a 7 bp inverted repeat (blue sequences highlighted with small arrows) separated by 11 nucleotides of non-complementary sequence, and various restriction sites to facilitate evaluation of the fork and its products. (B) The single stranded stretch on the lagging strand exposes two complementary copies of the inverted repeat (in blue with arrow) which are expected to fold into a stem-loop structure to achieve the lowest free energy state for the forked molecule. (C and D) Breathing of the duplexes at the fork facilitates fork reversal and generates alternative conformations by branch migration—all with identical numbers of paired bases. The structures in B and D reflect the two extreme branch-migrated forms, with C representing only one of 25 possible intermediates. The two nucleotides in red are brought into proximity in the branch-migrated forms, providing a substrate for ligation by DNA ligase. Completion of the ligation step creates an XhoI restriction site. PCR primers (shown in C) target the two linkers.