FIG. 6. Comparing modeled errors in measured pIC50 values using tip-based or acoustic direct dispensing.
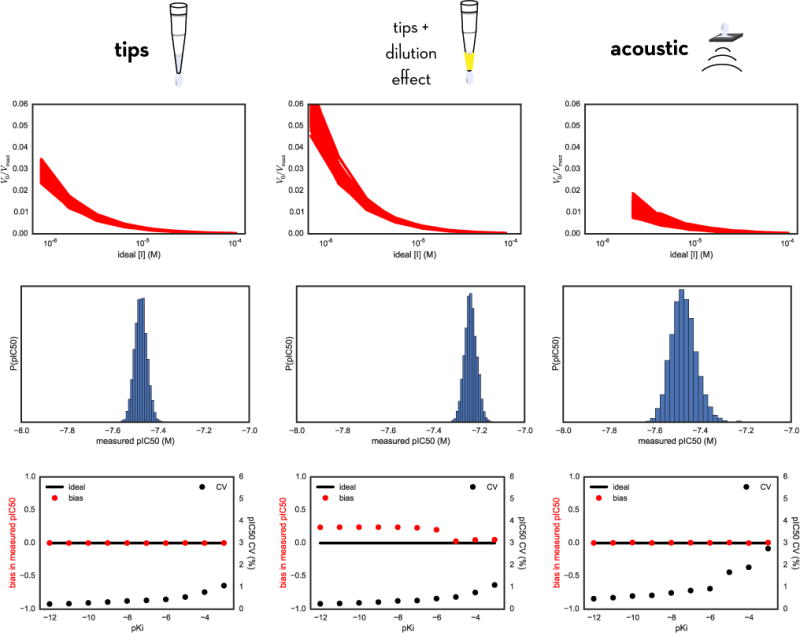
Top row: Bootstrap simulation of the entire assay yields a distribution of V0/Vmax (proportional to measured product accumulation) vs ideal inhibitor concentration [I] curves for many synthetic bootstrap replicates of the assay. Here, the inhibitor is modeled to have a true Ki of 1 nM (pKi = −9). Middle row: For the same inhibitor, we obtain a distribution of measured pIC50 values from fitting the Using our model we can look at the variance in activity measurements as a function of inhibitor concentration [I] (top), which then directly translates into a distribution of measured pIC50 values. Bottom row: Scaning across a range of true compound affinities, we can repeat the bootstrap sampling procedure and analyze the distribution of measured pIC50 values to obtain estimates of the relative bias (red) and CV (black) for the resulting measured pIC50s. For all methods, the CV increases for weaker affinities; for tip-based dispensing using fixed tips and incorporating the dilution effect, a significant bias is notable.
