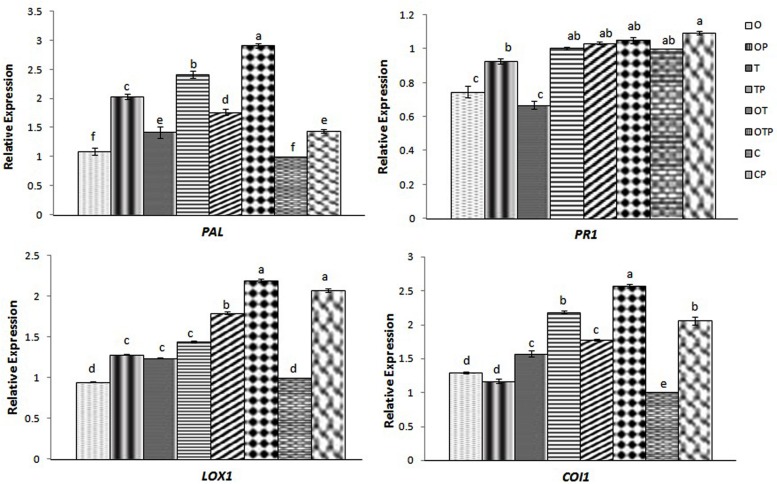
FIGURE 2.
Relative transcript accumulation patterns of two key genes representing JA (LOX1 and COI1) and SA (PAL and PR1) -mediated defense responses in pea leaves during challenge by the biotroph fungal pathogen Erysiphe as analyzed through quantitative real time-PCR in different treatments after 24 h of pathogen (E. pisi) challenge: O- P. fluorescens OKC; OP- OKC treated and pathogen challenged; T- T. asperellum T42; TP- T42 treated and pathogen challenged; OT- combination of OKC and T42; OTP- OKC and T42 treated and pathogen challenged; P- Pathogen challenged and C- OKC and T42 non-treated and pathogen unchallenged control. The qRT-PCR data were normalized with mean CT value of the endogenous gene ubiquitin and fold accumulation of transcripts was compared by using the mean of the CT values of the three biological replicates with treatment C (control). Error bars represent SD from means of three measurements. Different superscript letters indicate data significantly different from the other treatments (P ≤ 0.05; Duncan’s multiple range test).