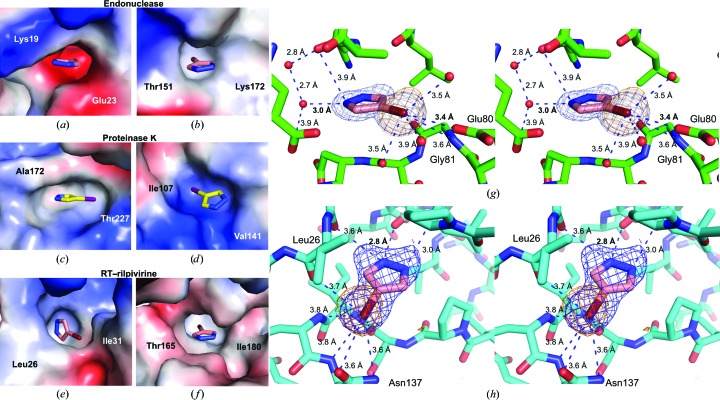
Figure 4.
4-Bromopyrazole and 4-iodopyrazole binding sites. (a)–(f) Two binding sites with the clearest electron density for the compounds are depicted for each protein: (a, b) influenza endonuclease, (c, d) proteinase K and (e, f) RT–rilpivirine. Electrostatic potential surfaces are shown. Two residues in each pocket are labeled. (g, h) Wall-eye stereoviews of 4-bromopyrazole bound to pockets in influenza endonuclease (g) or RT–rilpivirine (h). Potential electrostatic interactions are depicted as blue dashes with interatomic distances shown. Compound OMIT maps are contoured at 3.0σ (blue; F o − F c coefficients) and anomalous difference density is contoured at 5.0σ [orange mesh; F(hkl) − F(−h−k−l) coefficients].