Fig. 6.
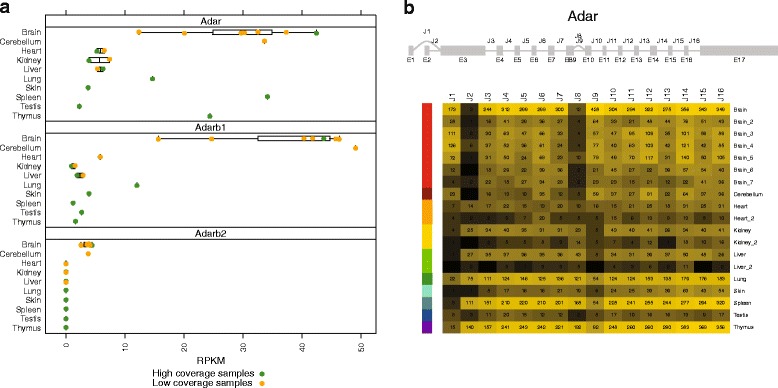
Expression level of Adar, Adarb1, and Adarb2 genes across tissues, as measured by RPKM (a). Splice junction usage across samples for ADAR transcripts (b). Raw counts of reads supporting each splice junction in each sample are included as text in the cells of the isoform heatmap. The heatmap color gradient represents the l o g2(F P K M+1) for each junction and scales from minimum (black) to maximum (yellow). ADAR is known to have a shorter, typically brain specific protein isoform (referred to as p110), and a more ubiquitously expressed longer protein isoform (p150). These two isoform variants are distinguished by the alternate 5’ UTRs that are used by the J1 and J2 mutually exclusive splicing events. J1 produces the shorter p110 isform while J2 produces the longer p150 isoform. Of note, an internal alternative splicing event, J8, tends to co-occur with the p150 isoform, while the J9 splice appears more frequently in samples with higher p110 isoform usage
