Fig. 3.
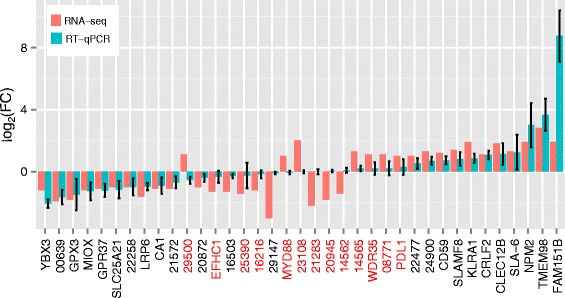
Validation of DEGs by RT-qPCR. 24 of the 37 selected DEGs between low and high RFI groups were confirmed by RT-qPCR when using the 48 samples, 24 of which were used for RNA-seq and another 24 of which were novel, as shown in this figure (q ≤ 0.15); 22 of the 37 selected DEGs were confirmed by RT-qPCR when using the 24 samples that were used for RNA-seq (q ≤ 0.15); but only 9 of the 37 selected DEGs were validated by RT-qPCR when using the 24 novel samples (q ≤ 0.15) (Addition file 9: Table S7). For comparison, the log2 (fold change) of these genes determined by RNA-seq were also displayed. Genes were ordered based on their log2 (fold change) as determined by RT-qPCR for display. Error bars for RT-qPCR assays show the standard errors of mean log2 (fold change). DEGs not confirmed by RT-qPCR are labeled in red. Genes without corresponding human orthologs are labeled with the last 5 digits of their Ensembl gene IDs, with common prefix “ENSSSCG000000” omitted for simplicity. For example, the ID for “29500” should be ENSSSCG00000029500
