Fig. 3.
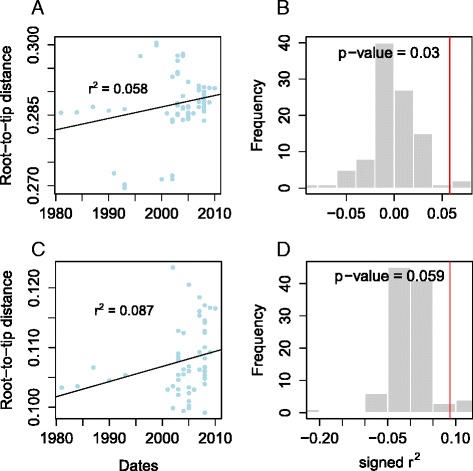
Investigating the presence of a clock-like signal in the data. The presence of a molecular clock in the dataset was assessed by plotting the root to tip distance of isolates in the phylogeny against isolation date [53] for both the dataset as a whole (a + b) and only Type II isolates (c + d). Both had evidence of a very weak positive signal, as indicated by a low linear regression correlation coefficient (a + c). In order to test the significance of these observations, 99 comparison datasets were produced, in which the isolation dates were permuted on the phylogeny. b and d show histograms of correlation coefficient values from the permuted datasets, with the red line indicating the value from the real data. In both cases the correlation coefficient of the real data is significantly greater than the permuted data at the 0.05 level. Passing this test is a minimal requirement for the application of BEAST analysis (Fig. 4), which assumes the presence of a molecular clock
