Figure 1.
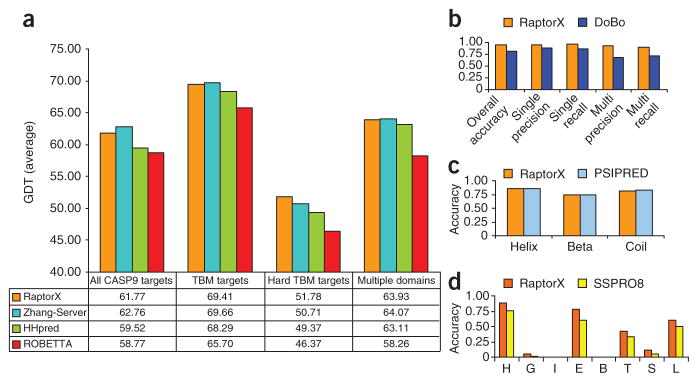
Performance assessment of core prediction modules in the RaptorX server. (a) Comparison of structure prediction performance by global distance test (GDT) score for RaptorX and three other publicly available protocols on the CASP9 targets. Performance is compared in four categories: All CASP9 targets, template-based modeling (TBM) targets, hard TBM targets and multidomain targets. (b) Performance comparison for domain parsing between RaptorX and DoBo. Metrics are given for overall performance, and performance on single-domain and multidomain CASP9 target proteins. Specifically, accuracy is the overall proportion of both single-domain and multidomain proteins identified correctly; single (multi) recall is the fraction of single-domain (multidomain) proteins that are predicted; single (multi) precision is the fraction of correctly predicted single-domain (multidomain) proteins among all the predictions. A multidomain protein is correctly predicted only if its domain boundaries are correctly identified. (c) Performance comparison between RaptorX and PSIPRED for secondary structure prediction. The accuracies achieved for three-state prediction (helix, sheet and coil) are compared. (d) Performance comparison between RaptorX and SSPRO8 for eight-state secondary structure prediction. The accuracies achieved for eight-state prediction for the classes H, G, I, E, B, T, S, L (using SSPRO8 nomenclature) are compared.