Abstract
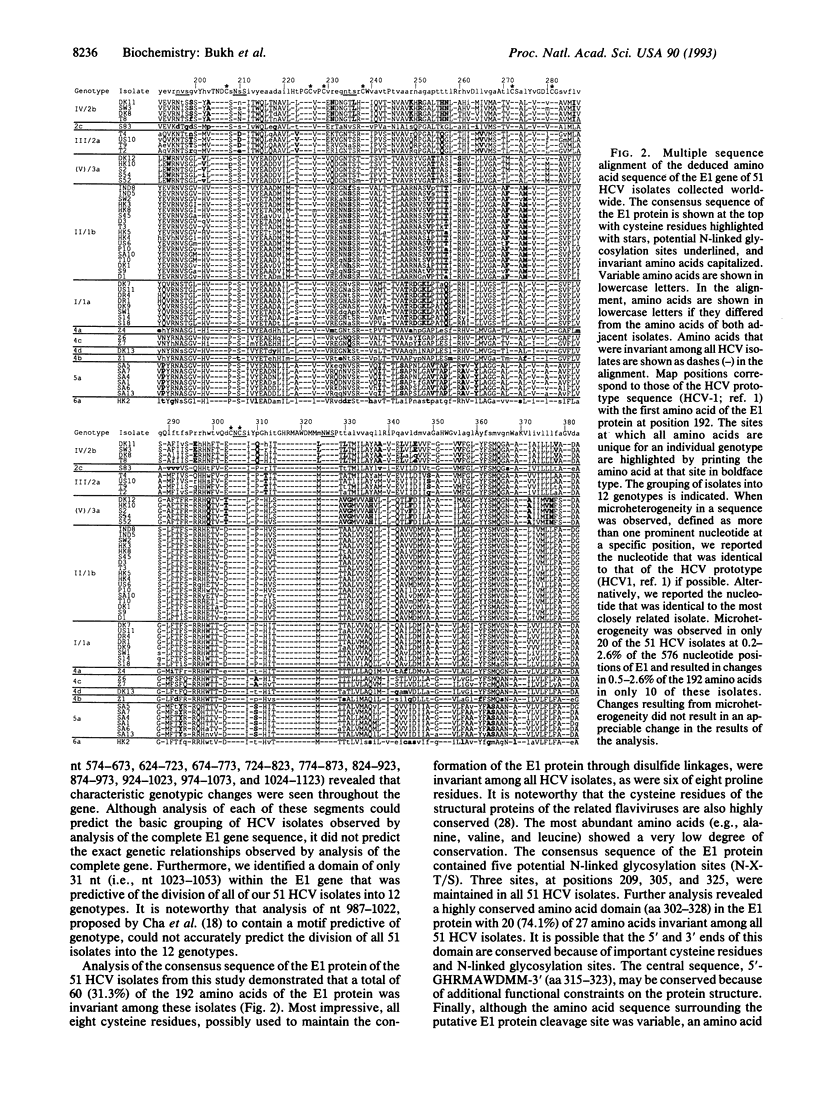
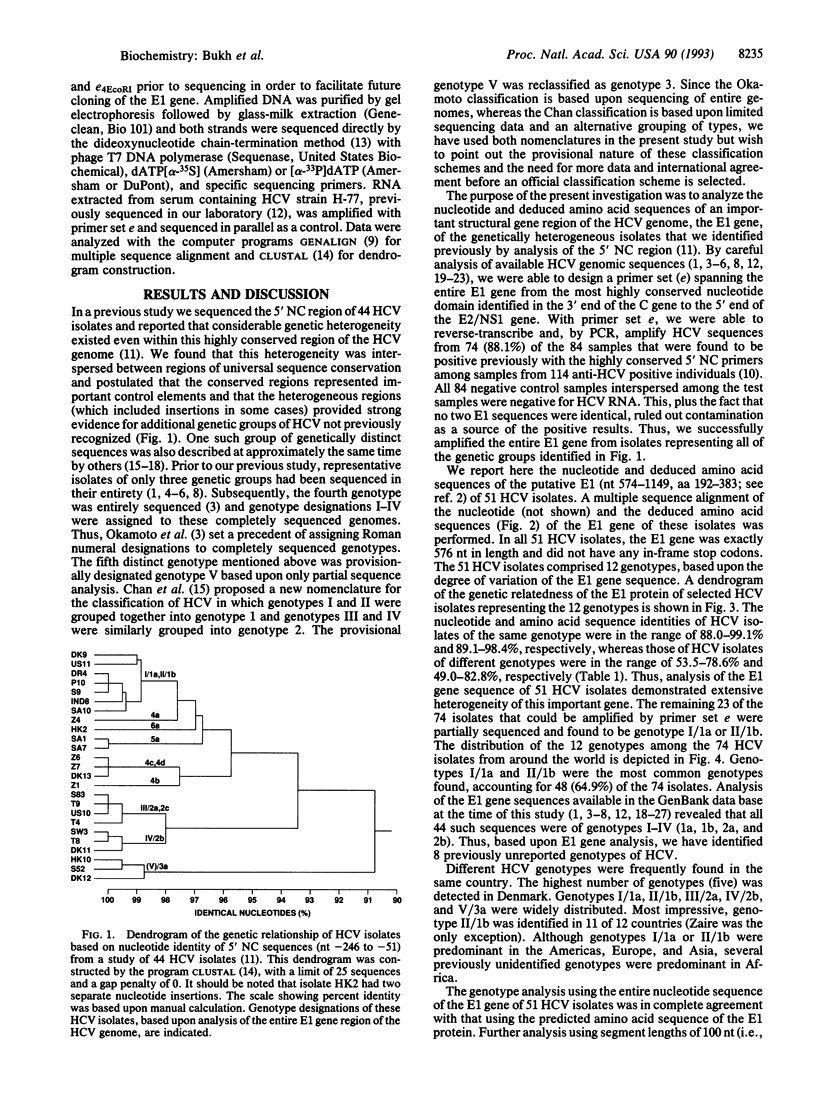
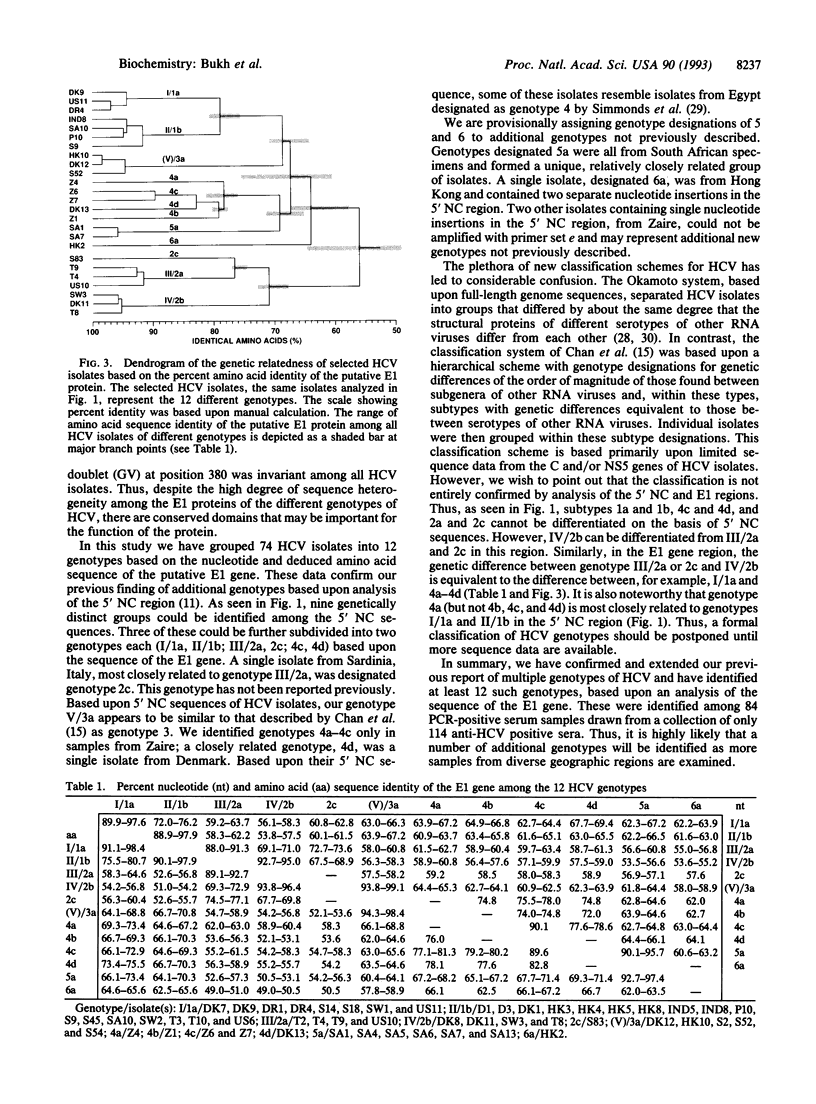
In a previous study we sequenced the 5' noncoding (NC) region of 44 isolates of hepatitis C virus (HCV) and identified heterogeneous domains that provided evidence for additional genetic groups of HCV not previously recognized. In this study we have determined the complete nucleotide sequence of the putative envelope 1 (E1) gene in 51 HCV isolates from around the world and found that they could be grouped into at least 12 distinct genotypes. The E1 gene sequence of 8 of these genotypes has not been reported previously. Although the genetic relatedness of HCV isolates determined by the previous analysis of the 5' NC region predicted the relationships observed in the E1 gene, analysis of the 5' NC sequence alone did not accurately predict all HCV genotypes. The nucleotide and amino acid sequence identities of the E1 gene among HCV isolates of the same genotype were in the range of 88.0-99.1% and 89.1-98.4%, respectively, whereas those of HCV isolates of different genotypes were in the range of 53.5-78.6% and 49.0-82.8%, respectively. The latter differences are similar to those found when comparing the envelope gene sequences of the various serotypes of the related flaviviruses as well as other RNA viruses. We found that some genotypes of HCV were widely distributed around the world, whereas others were identified only in discreet geographical regions. Four genotypes were identified exclusively in Africa and comprised the majority of HCV isolates on that continent. The E1 gene was exactly 576 nucleotides in length in all 51 HCV isolates with no in-frame stop codons. Analysis of the predicted E1 protein identified several conserved domains that may be important for maintaining its biological function: (i) eight invariant cysteine residues, (ii) three potential N-linked glycosylation sites, (iii) a domain of nine amino acids (GHRMAWDMM), and (iv) an amino acid doublet (GV) near the putative cleavage site at the C terminus of the protein. In conclusion, the discovery of at least 12 genotypes of HCV has important implications for HCV diagnosis and vaccine development.
Full text
PDF


Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Abe K., Inchauspe G., Fujisawa K. Genomic characterization and mutation rate of hepatitis C virus isolated from a patient who contracted hepatitis during an epidemic of non-A, non-B hepatitis in Japan. J Gen Virol. 1992 Oct;73(Pt 10):2725–2729. doi: 10.1099/0022-1317-73-10-2725. [DOI] [PubMed] [Google Scholar]
- Bachmann B., Lüke W., Hunsmann G. Improvement of PCR amplified DNA sequencing with the aid of detergents. Nucleic Acids Res. 1990 Mar 11;18(5):1309–1309. doi: 10.1093/nar/18.5.1309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bukh J., Purcell R. H., Miller R. H. Importance of primer selection for the detection of hepatitis C virus RNA with the polymerase chain reaction assay. Proc Natl Acad Sci U S A. 1992 Jan 1;89(1):187–191. doi: 10.1073/pnas.89.1.187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bukh J., Purcell R. H., Miller R. H. Sequence analysis of the 5' noncoding region of hepatitis C virus. Proc Natl Acad Sci U S A. 1992 Jun 1;89(11):4942–4946. doi: 10.1073/pnas.89.11.4942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cha T. A., Beall E., Irvine B., Kolberg J., Chien D., Kuo G., Urdea M. S. At least five related, but distinct, hepatitis C viral genotypes exist. Proc Natl Acad Sci U S A. 1992 Aug 1;89(15):7144–7148. doi: 10.1073/pnas.89.15.7144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chan S. W., McOmish F., Holmes E. C., Dow B., Peutherer J. F., Follett E., Yap P. L., Simmonds P. Analysis of a new hepatitis C virus type and its phylogenetic relationship to existing variants. J Gen Virol. 1992 May;73(Pt 5):1131–1141. doi: 10.1099/0022-1317-73-5-1131. [DOI] [PubMed] [Google Scholar]
- Chen P. J., Lin M. H., Tai K. F., Liu P. C., Lin C. J., Chen D. S. The Taiwanese hepatitis C virus genome: sequence determination and mapping the 5' termini of viral genomic and antigenomic RNA. Virology. 1992 May;188(1):102–113. doi: 10.1016/0042-6822(92)90739-c. [DOI] [PubMed] [Google Scholar]
- Choo Q. L., Richman K. H., Han J. H., Berger K., Lee C., Dong C., Gallegos C., Coit D., Medina-Selby R., Barr P. J. Genetic organization and diversity of the hepatitis C virus. Proc Natl Acad Sci U S A. 1991 Mar 15;88(6):2451–2455. doi: 10.1073/pnas.88.6.2451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hahn Y. S., Galler R., Hunkapiller T., Dalrymple J. M., Strauss J. H., Strauss E. G. Nucleotide sequence of dengue 2 RNA and comparison of the encoded proteins with those of other flaviviruses. Virology. 1988 Jan;162(1):167–180. doi: 10.1016/0042-6822(88)90406-0. [DOI] [PubMed] [Google Scholar]
- Higgins D. G., Sharp P. M. CLUSTAL: a package for performing multiple sequence alignment on a microcomputer. Gene. 1988 Dec 15;73(1):237–244. doi: 10.1016/0378-1119(88)90330-7. [DOI] [PubMed] [Google Scholar]
- Hijikata M., Kato N., Ootsuyama Y., Nakagawa M., Ohkoshi S., Shimotohno K. Hypervariable regions in the putative glycoprotein of hepatitis C virus. Biochem Biophys Res Commun. 1991 Feb 28;175(1):220–228. doi: 10.1016/s0006-291x(05)81223-9. [DOI] [PubMed] [Google Scholar]
- Honda M., Kaneko S., Unoura M., Kobayashi K., Murakami S. Sequence analysis of putative structural regions of hepatitis C virus isolated from 5 Japanese patients with hepatocellular carcinoma. Arch Virol. 1993;128(1-2):163–169. doi: 10.1007/BF01309797. [DOI] [PubMed] [Google Scholar]
- Houghton M., Weiner A., Han J., Kuo G., Choo Q. L. Molecular biology of the hepatitis C viruses: implications for diagnosis, development and control of viral disease. Hepatology. 1991 Aug;14(2):381–388. [PubMed] [Google Scholar]
- Inchauspe G., Zebedee S., Lee D. H., Sugitani M., Nasoff M., Prince A. M. Genomic structure of the human prototype strain H of hepatitis C virus: comparison with American and Japanese isolates. Proc Natl Acad Sci U S A. 1991 Nov 15;88(22):10292–10296. doi: 10.1073/pnas.88.22.10292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kato N., Hijikata M., Ootsuyama Y., Nakagawa M., Ohkoshi S., Sugimura T., Shimotohno K. Molecular cloning of the human hepatitis C virus genome from Japanese patients with non-A, non-B hepatitis. Proc Natl Acad Sci U S A. 1990 Dec;87(24):9524–9528. doi: 10.1073/pnas.87.24.9524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee C. H., Cheng C., Wang J., Lumeng L. Identification of hepatitis C viruses with a nonconserved sequence of the 5' untranslated region. J Clin Microbiol. 1992 Jun;30(6):1602–1604. doi: 10.1128/jcm.30.6.1602-1604.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu K., Hu Z., Li H., Prince A. M., Inchauspe G. Genomic typing of hepatitis C viruses present in China. Gene. 1992 May 15;114(2):245–250. doi: 10.1016/0378-1119(92)90582-a. [DOI] [PubMed] [Google Scholar]
- Miller R. H., Purcell R. H. Hepatitis C virus shares amino acid sequence similarity with pestiviruses and flaviviruses as well as members of two plant virus supergroups. Proc Natl Acad Sci U S A. 1990 Mar;87(6):2057–2061. doi: 10.1073/pnas.87.6.2057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mori S., Kato N., Yagyu A., Tanaka T., Ikeda Y., Petchclai B., Chiewsilp P., Kurimura T., Shimotohno K. A new type of hepatitis C virus in patients in Thailand. Biochem Biophys Res Commun. 1992 Feb 28;183(1):334–342. doi: 10.1016/0006-291x(92)91648-a. [DOI] [PubMed] [Google Scholar]
- Ogata N., Alter H. J., Miller R. H., Purcell R. H. Nucleotide sequence and mutation rate of the H strain of hepatitis C virus. Proc Natl Acad Sci U S A. 1991 Apr 15;88(8):3392–3396. doi: 10.1073/pnas.88.8.3392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Okamoto H., Kojima M., Okada S., Yoshizawa H., Iizuka H., Tanaka T., Muchmore E. E., Peterson D. A., Ito Y., Mishiro S. Genetic drift of hepatitis C virus during an 8.2-year infection in a chimpanzee: variability and stability. Virology. 1992 Oct;190(2):894–899. doi: 10.1016/0042-6822(92)90933-g. [DOI] [PubMed] [Google Scholar]
- Okamoto H., Kurai K., Okada S., Yamamoto K., Lizuka H., Tanaka T., Fukuda S., Tsuda F., Mishiro S. Full-length sequence of a hepatitis C virus genome having poor homology to reported isolates: comparative study of four distinct genotypes. Virology. 1992 May;188(1):331–341. doi: 10.1016/0042-6822(92)90762-e. [DOI] [PubMed] [Google Scholar]
- Okamoto H., Okada S., Sugiyama Y., Kurai K., Iizuka H., Machida A., Miyakawa Y., Mayumi M. Nucleotide sequence of the genomic RNA of hepatitis C virus isolated from a human carrier: comparison with reported isolates for conserved and divergent regions. J Gen Virol. 1991 Nov;72(Pt 11):2697–2704. doi: 10.1099/0022-1317-72-11-2697. [DOI] [PubMed] [Google Scholar]
- Okamoto H., Okada S., Sugiyama Y., Yotsumoto S., Tanaka T., Yoshizawa H., Tsuda F., Miyakawa Y., Mayumi M. The 5'-terminal sequence of the hepatitis C virus genome. Jpn J Exp Med. 1990 Jun;60(3):167–177. [PubMed] [Google Scholar]
- Simmonds P., McOmish F., Yap P. L., Chan S. W., Lin C. K., Dusheiko G., Saeed A. A., Holmes E. C. Sequence variability in the 5' non-coding region of hepatitis C virus: identification of a new virus type and restrictions on sequence diversity. J Gen Virol. 1993 Apr;74(Pt 4):661–668. doi: 10.1099/0022-1317-74-4-661. [DOI] [PubMed] [Google Scholar]
- Takamizawa A., Mori C., Fuke I., Manabe S., Murakami S., Fujita J., Onishi E., Andoh T., Yoshida I., Okayama H. Structure and organization of the hepatitis C virus genome isolated from human carriers. J Virol. 1991 Mar;65(3):1105–1113. doi: 10.1128/jvi.65.3.1105-1113.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takeuchi K., Kubo Y., Boonmar S., Watanabe Y., Katayama T., Choo Q. L., Kuo G., Houghton M., Saito I., Miyamura T. The putative nucleocapsid and envelope protein genes of hepatitis C virus determined by comparison of the nucleotide sequences of two isolates derived from an experimentally infected chimpanzee and healthy human carriers. J Gen Virol. 1990 Dec;71(Pt 12):3027–3033. doi: 10.1099/0022-1317-71-12-3027. [DOI] [PubMed] [Google Scholar]
- Tanaka T., Kato N., Nakagawa M., Ootsuyama Y., Cho M. J., Nakazawa T., Hijikata M., Ishimura Y., Shimotohno K. Molecular cloning of hepatitis C virus genome from a single Japanese carrier: sequence variation within the same individual and among infected individuals. Virus Res. 1992 Apr;23(1-2):39–53. doi: 10.1016/0168-1702(92)90066-i. [DOI] [PubMed] [Google Scholar]
- Weiner A. J., Brauer M. J., Rosenblatt J., Richman K. H., Tung J., Crawford K., Bonino F., Saracco G., Choo Q. L., Houghton M. Variable and hypervariable domains are found in the regions of HCV corresponding to the flavivirus envelope and NS1 proteins and the pestivirus envelope glycoproteins. Virology. 1991 Feb;180(2):842–848. doi: 10.1016/0042-6822(91)90104-j. [DOI] [PubMed] [Google Scholar]


