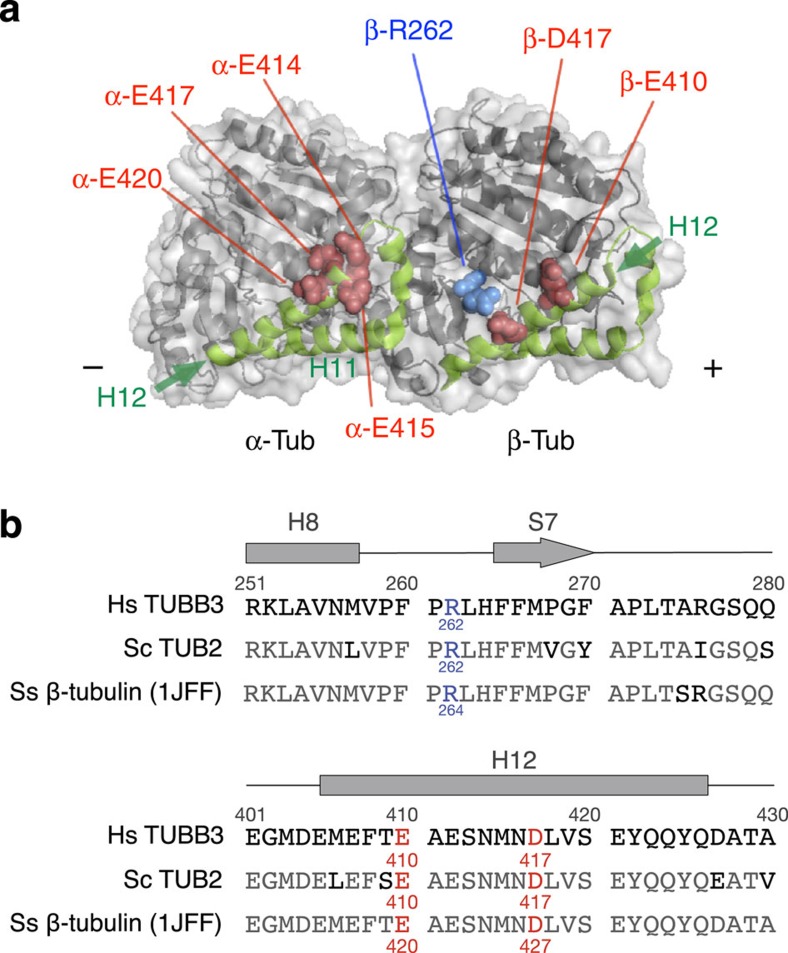
Figure 1. The position of residue β-R262 in the structure of tubulin dimer and in its amino-acid sequence.
(a) Distribution of the residues involved in the interaction with kinesin on tubulin dimer (on face view of the MT; PDB: 1JFF). Alanine-scanning mutagenesis analysis of tubulin identified six acidic residues (red spheres) in the C-terminal hairpin structures (H11–H11′–H12) of α- and β-tubulin (green) as critical for the interaction with kinesin17. The basic residue β-R262 addressed in this paper (blue spheres) is located in the H8-S7 loop of β-tubulin. (b) Alignment of the sequences of human (Hs) TUBB3, Saccharomyces cerevisiae (Sc) TUB2, and pig (Ss) β-tubulin. For pig tubulin, sequence numbering used in the PDB 1JFF was adopted. The positions of the secondary structure elements are indicated, and the labelled residues correspond to those shown in a with the same colour scheme.