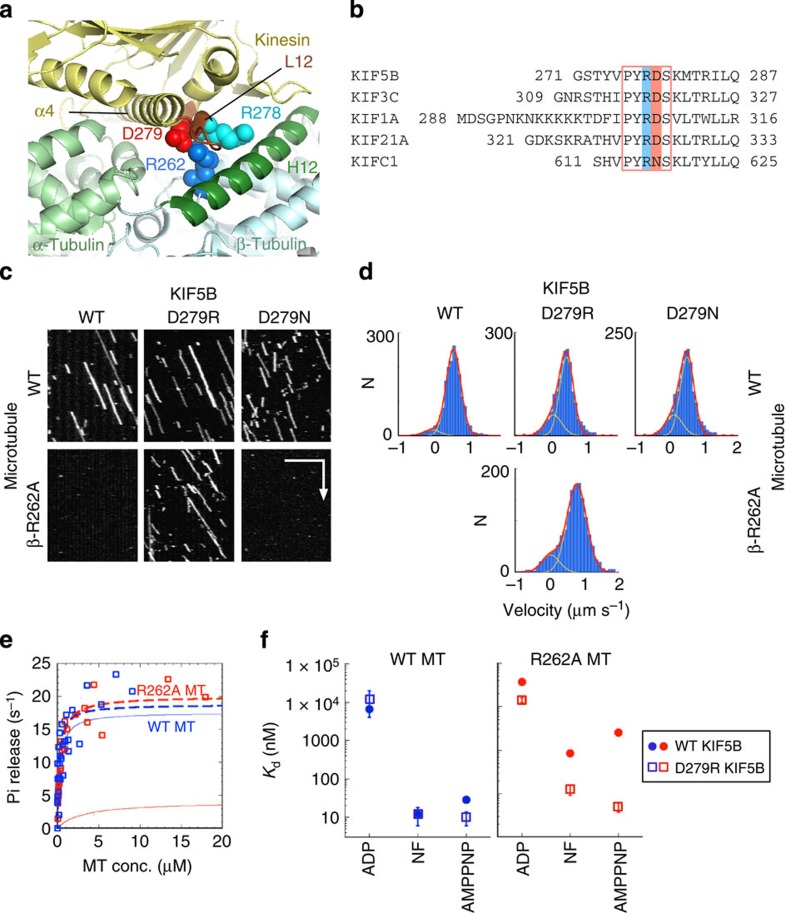
Figure 3. Motility rescue by D279R KIF5B.
(a) Molecular structure of the kinesin–MT interface surrounding β-R262 of tubulin (PDB: 4LNU)20. A pair of charged residues, R278 (cyan spheres) and D279 (red spheres), in the kinesin L12 loop (brown) can be seen facing R262 in β-tubulin (blue spheres). (b) Alignment of L12 loop and the adjacent α5 helix sequences. The PYRDS motif (red box) is highly conserved in the kinesin families. (c) Kymographs of KIF5B motility on WT MTs and β-R262A MTs. Horizontal bar, 5 μm; vertical arrow, 5 s. Motility parameters are given in Table 1. (d) Histogram of the KIF5B velocities on WT and β-R262A MTs. (e) The ATPase activity of D279R KIF5B activated by WT MTs (□) or R262A MTs (□). All data obtained from four and three independent measurements (for WT and β-R262A, respectively) are plotted in the graph. Dashed curves represent the best fits to the Michaelis–Menten equation with kcat and K0.5MT values given in Table 1. The ATPase activities of WT KIF5B (shown in Fig. 2b) are included as references (—, WT–MT; —, R262A MT). (f) Dissociation constants of the KIF5B–MT complex determined by a co-sedimentation assay. ‘NF' represents a nucleotide-free condition. Errors indicate errors in fitting the curve of the equilibrium binding data to the hyperbola (the raw data and exact number of Kd values are shown in Supplementary Table 2).