Fig. 1.
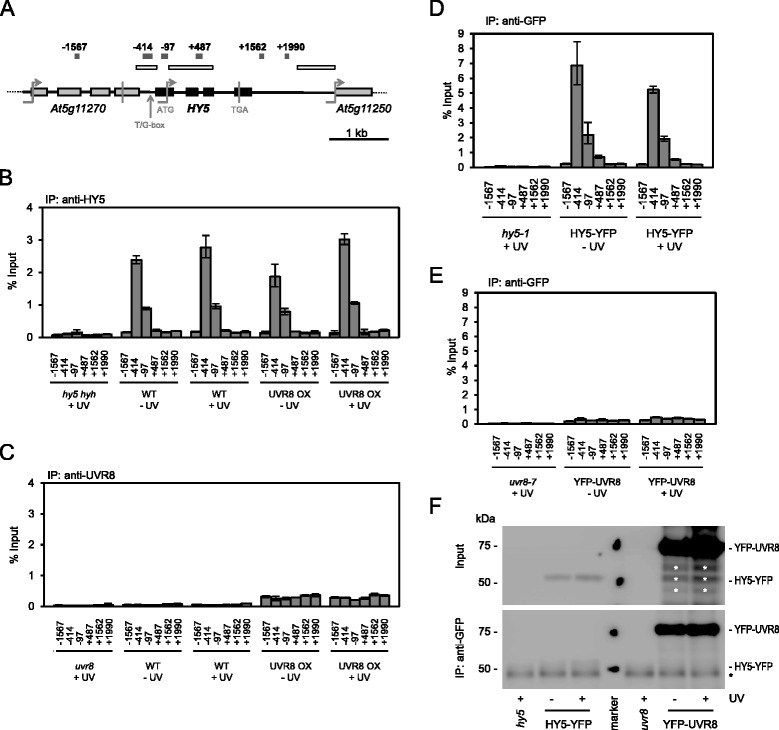
UVR8 does not associate with the HY5 genomic region. a Schematic representation of the HY5 genomic locus and surrounding region. Black and grey boxes depict the HY5 transcribed region (with start ATG and stop TGA of HY5 indicated) and portions of the transcribed regions of the neighbouring genes, respectively. Upper grey bars indicate qRT-PCR amplicons at various positions at and surrounding the HY5 genomic locus with numbers indicating the position of the 5’ end of the amplicon relative to the HY5 translation start site (referred to as position +1). An arrow indicates the position of the T/G-box cis-element bound by HY5 [11, 12]. Open bars are placed at genomic regions previously reported to be UVR8 binding targets in gel-based ChIP assays [19]. b, c ChIP-qPCR of DNA associated with HY5 (b) and UVR8 (c) presented as the percentage of total input DNA recovered by immunoprecipitation (% Input). The same chromatin pool was used for immunoprecipitation with both HY5 and UVR8 antibodies. WT = Ws/Pro HY5 :Luc+, UVR8 OX = Ws/Pro HY5 :Luc + Pro 35S :UVR8. Arabidopsis mutant lines with corresponding genetic backgrounds (uvr8 = uvr8-7/Pro HY5 :Luc+, hy5 hyh = hy5-ks50 hyh-1/Pro HY5 :Luc+) were included as negative controls. d, e As in (b, c), but of DNA associated with HY5-YFP (d) and YFP-UVR8 (e), performed using the same anti-GFP antibody for immunoprecipitation. HY5-YFP = hy5-1/Pro 35S :HY5-YFP, YFP-UVR8 = uvr8-7/Pro 35S :YFP-UVR8. Arabidopsis mutant lines with corresponding genetic backgrounds (hy5-1 and uvr8-7) were included as negative controls. b, c, d, e The number labelling indicates the location of each analysed DNA fragment, specifically the position of the 5’ end of the amplicon relative to the translation start site (referred to as position +1). Experimental material was sourced from ten-day-old seedlings, either processed with no UV-B exposure (−UV) or following 4 h UV-B (+UV). Error bars represent SDs of three technical replicates. f Input and eluate of D/E ChIP samples were analysed on western blots with an antibody against GFP. Bands resulting from degradation products (upper panel) and unspecific background signal (lower panel) are marked (*). Molecular weight marker positions representing 50 and 75 kDa are shown (marker lane)
