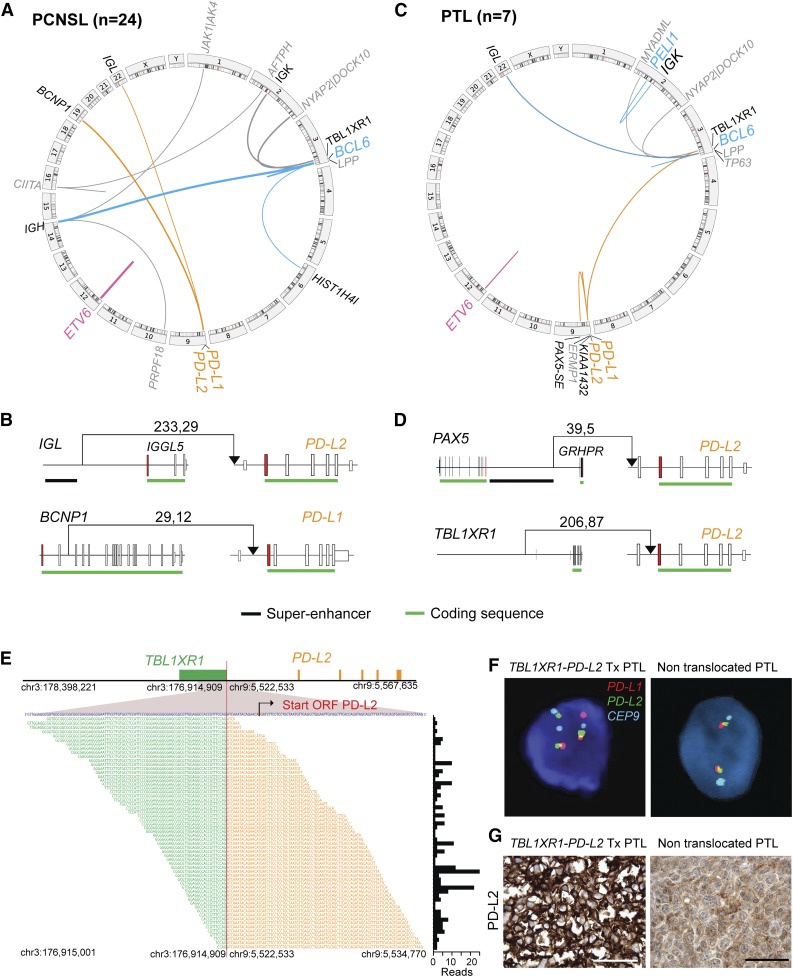
Figure 3.
Chromosomal rearrangements in PCNSL and PTL. (A,C) Detected chromosomal rearrangements in 24 PCNSL (A) and 7 PTL (C) are summarized as circos plots. Structural alterations involving certain partners are highlighted; BCL6, blue; ETV6, pink; PD-1 ligands, orange. Partners of color-coded alterations are black, all other alterations are gray. Frequency of events is indicated by line thickness. (B,D) Chromosomal rearrangements involving PD-L1 or PD-L2 are plotted in their genomic context. Exons are visualized as boxes, ATG-containing exon are in red, the coding region is underlined in green, and previously identified super-enhancers in DLBCLs43 are underlined in black. The number of supporting reads (split reads, read pairs) is indicated above each translocation. (E) TBL1XR1-PD-L2 fusion as validated by RNA-Seq. Chromosomal breakpoint is depicted by the red line. Start codon of PD-L2 is indicated in red within the contig of the RNA-Seq reads. The translocation involves only the regulatory elements of TBL1XR1 and does not affect the open reading frame (ORF) of PD-L2. Individual supporting reads are shown in the lower panel, with frequencies as a bar graph on the right. (F) FISH assays of PTLs with the PD-L2 translocation (left panel) or with wild-type PD-L2 (right panel). PD-L1 in red, PD-L2 in green, and centromeric probe (CEP9) in aqua. (G) IHC of PD-L2 expression in the translocated PTL (left panel) and a PTL with wild-type PD-L2 (right panel). The scale bar represents 100 μm.