Fig. 3.
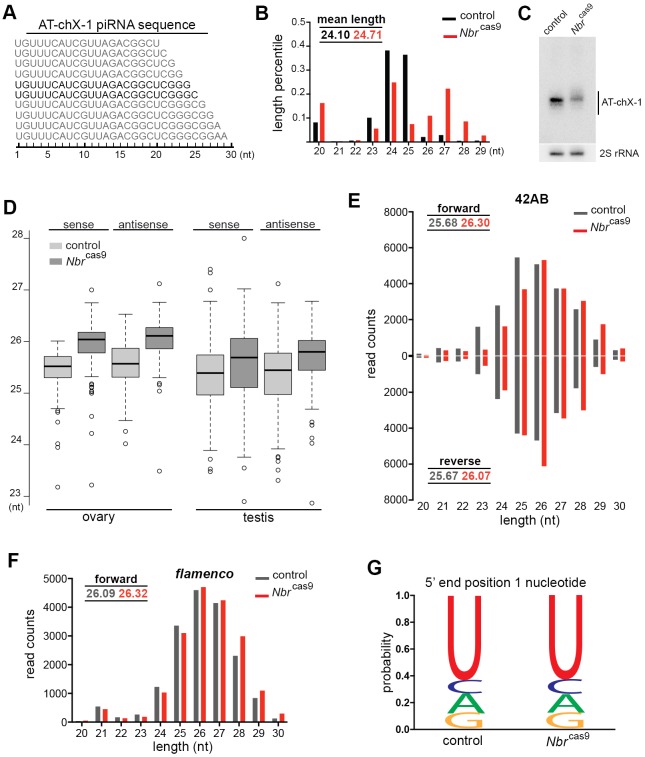
Nbr trims piRNAs at the 3′ end. (A) AT-chX-1, a piRNA enriched in the testis, displays the sequence feature of a 5′ uridine and a nested series at the 3′ end. The main forms are 24 and 25 nt (bold). (B) Length distribution analysis indicates that AT-chX-1 accumulates more long forms in Nbrcas9 (red) than in the control (black). RNAs were from testis. Genotypes: control (5905) and Nbrcas9/cas9 (Nbrcas9). (C) Small RNA northern blot confirms that AT-chX-1 shows a striking size increase in length in Nbrcas9 compared with control. 2S rRNA was used as a loading control. RNAs were from testis. (D) Gonadal piRNAs become longer in Nbr mutants than in controls. Box plots for length distribution revealed that Nbrcas9 flies accumulated more piRNAs with long forms than in controls. Using piPipes algorithms, only piRNAs mapped to transposons were used for subsequent analysis. Each transposon is assigned a mean value based on the length of mapped piRNAs, with a total of 127 individual transposons analyzed. Control versus Nbrcas9: ovary sense, P<2.2×10−16; ovary antisense, P<2.2×10−16; testis sense, P=4.861×10−7; testis antisense, P=1.19×10−10; Wilcoxon signed-rank test. RNAs were from ovaries and testis. (E,F) In the 42AB cluster (E), piRNAs can be derived from both forward (top) and reverse (bottom) strands, and they show accumulation of long forms regardless of their strand origin. In the flamenco cluster (F), piRNAs can only be generated from the forward strand (top), and they show Nbr-dependent length modulation. Mean lengths for piRNAs derived from the indicated clusters in control and Nbrcas9 are shown. RNAs were from ovaries. (G) Antisense piRNAs in Nbrcas9 show the same bias for 5′ uridine (1U) as in the control. (C-G) Genotypes as in B.