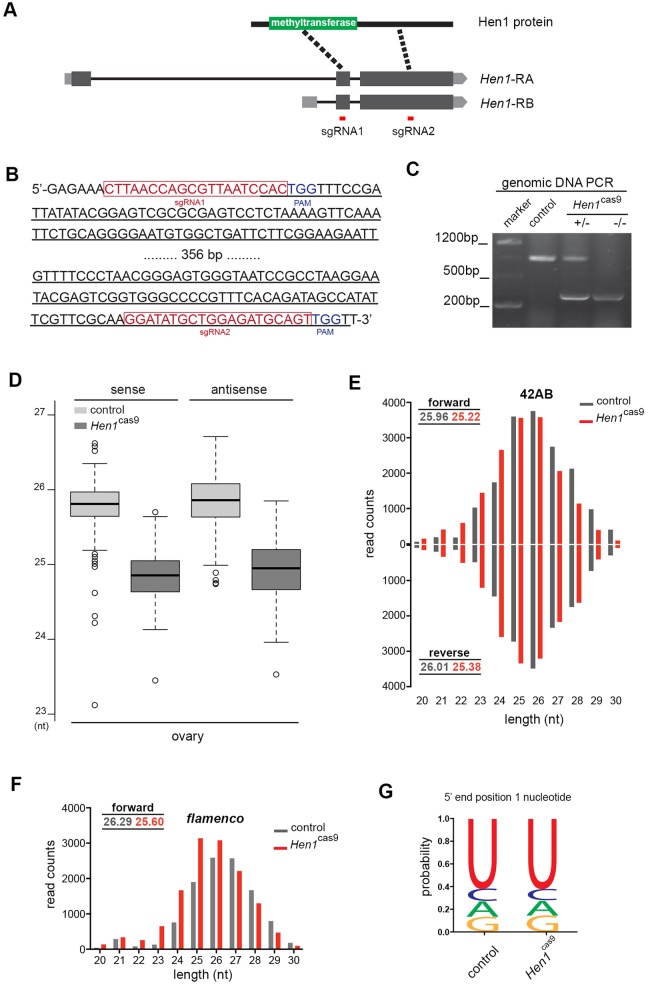
Fig. 4.
Loss of Hen1 shortens piRNAs from the 3′ end. (A) The Hen1 locus and guiding RNAs used to make Hen1cas9. To produce the Hen1 loss-of-function allele, sgRNA1 and sgRNA2 (red lines) were used to induce site-specific deletion. The deleted region largely removed the methyltransferase domain. (B) Part of the Hen1 genomic sequence illustrating features of the CRISPR/Cas9-mediated Hen1cas9 loss-of-function allele. sgRNAs and PAMs are highlighted. Resulting Hen1 mutant flies were backcrossed to the control homogeneous background for five generations to ensure background clearance. (C) PCR analysis confirms that Hen1cas9 is a deletion allele. DNAs were from whole flies. Genotypes: control (5905), Hen1cas9/+ and Hen1cas9/cas9. (D) Box plots for length distribution reveal that Hen1cas9 flies accumulate more short form piRNAs than controls. Control versus Hen1cas9: ovary sense, P<2.2×10−16; ovary antisense, P<2.2×10−16; Wilcoxon signed-rank test. RNAs were from ovaries. Genotypes: control (5905) and Hen1cas9/cas9 (Hen1cas9). (E,F) In the 42AB cluster (E) and the flamenco cluster (F), piRNAs show accumulation of shorter forms. Mean lengths for piRNAs derived from the indicated clusters in control and Hen1cas9 are shown. RNAs were from ovaries. (G) Antisense piRNAs in Hen1cas9 show the same bias for 5′ uridine (1U) as in control. RNAs were from ovaries. (E-G) Genotypes as in D.